All in One View
Content from Introduction to R and RStudio
Last updated on 2026-04-21 | Edit this page
Estimated time: 55 minutes
Overview
Questions
- How to find your way around RStudio?
- How to interact with R?
- How to manage your environment?
- How to install packages?
Objectives
- Describe the purpose and use of each pane in RStudio
- Locate buttons and options in RStudio
- Define a variable
- Assign data to a variable
- Manage a workspace in an interactive R session
- Use mathematical and comparison operators
- Call functions
- Manage packages
Before Starting The Workshop
Please ensure you have the latest version of R and RStudio installed on your machine. This is important, as some packages used in the workshop may not install correctly (or at all) if R is not up to date.
Why use R and R studio?
Welcome to the R portion of the Software Carpentry workshop!
Science is a multi-step process: once you’ve designed an experiment and collected data, the real fun begins with analysis! Throughout this lesson, we’re going to teach you some of the fundamentals of the R language as well as some best practices for organizing code for scientific projects that will make your life easier.
Although we could use a spreadsheet in Microsoft Excel or Google sheets to analyze our data, these tools are limited in their flexibility and accessibility. Critically, they also are difficult to share steps which explore and change the raw data, which is key to “reproducible” research.
Therefore, this lesson will teach you how to begin exploring your data using R and RStudio. The R program is available for Windows, Mac, and Linux operating systems, and is a freely-available where you downloaded it above. To run R, all you need is the R program.
However, to make using R easier, we will use the program RStudio, which we also downloaded above. RStudio is a free, open-source, Integrated Development Environment (IDE). It provides a built-in editor, works on all platforms (including on servers) and provides many advantages such as integration with version control and project management.
Overview
We will begin with raw data, perform exploratory analyses, and learn how to plot results graphically. This example starts with a dataset from gapminder.org containing population information for many countries through time. Can you read the data into R? Can you plot the population for Senegal? Can you calculate the average income for countries on the continent of Asia? By the end of these lessons you will be able to do things like plot the populations for all of these countries in under a minute!
Basic layout
When you first open RStudio, you will be greeted by three panels:
- The interactive R console/Terminal (entire left)
- Environment/History/Connections (tabbed in upper right)
- Files/Plots/Packages/Help/Viewer (tabbed in lower right)
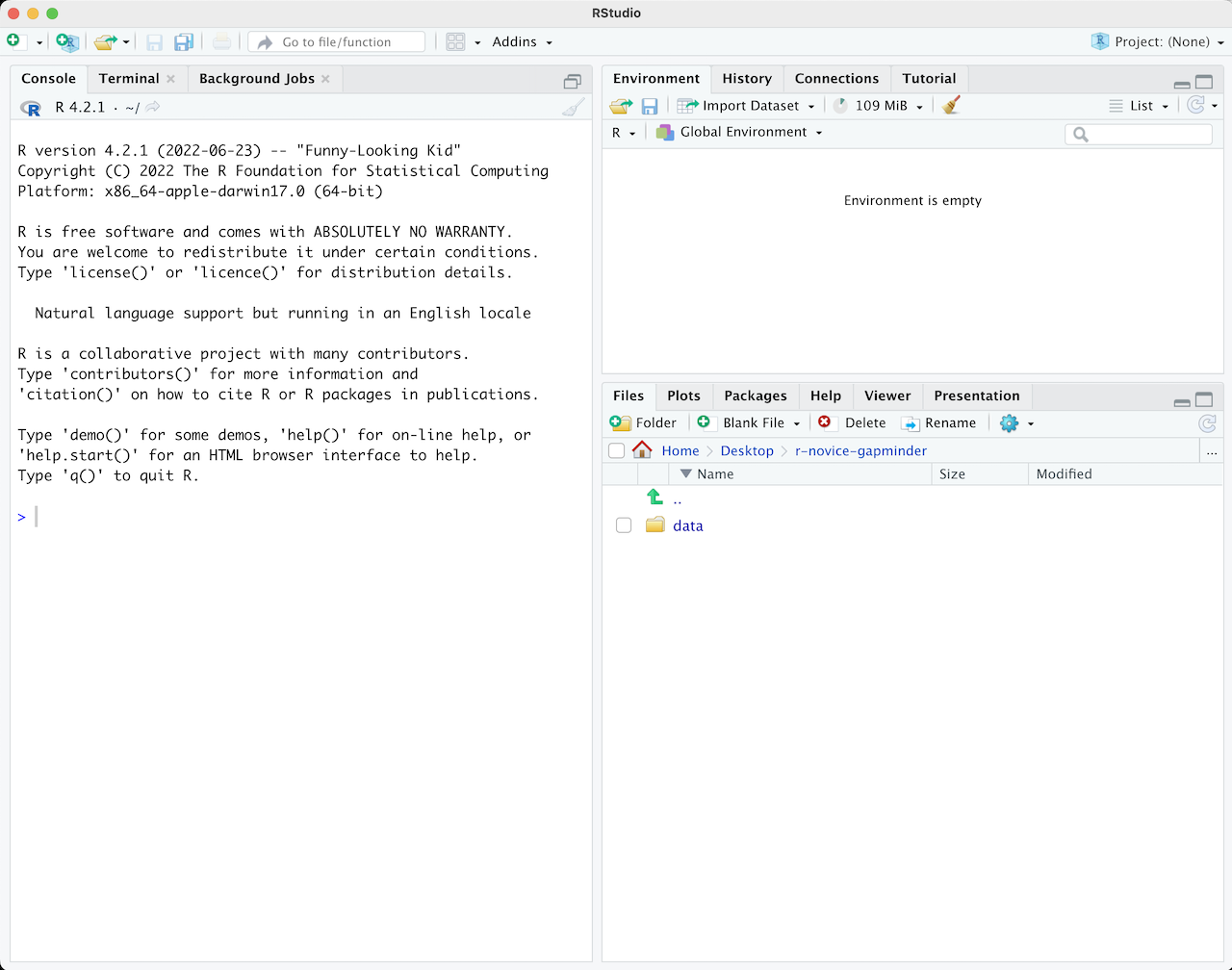
Once you open files, such as R scripts, an editor panel will also open in the top left.
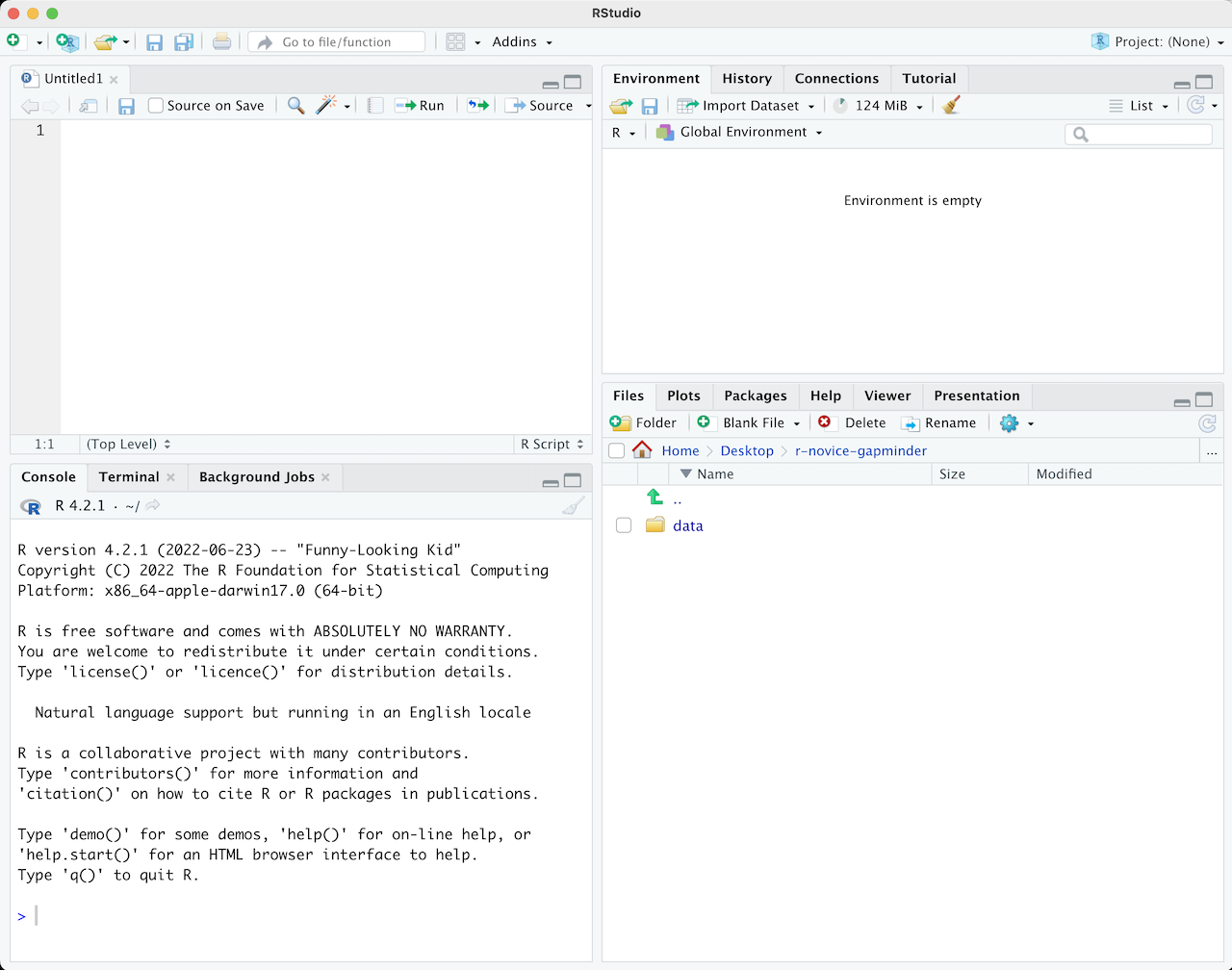
R scripts
Any commands that you write in the R console can be saved to a file
to be re-run again. Files containing R code to be ran in this way are
called R scripts. R scripts have .R at the end of their
names to let you know what they are.
Workflow within RStudio
There are two main ways one can work within RStudio:
- Test and play within the interactive R console then copy code into a .R file to run later.
- This works well when doing small tests and initially starting off.
- It quickly becomes laborious
- Start writing in a .R file and use RStudio’s short cut keys for the Run command to push the current line, selected lines or modified lines to the interactive R console.
- This is a great way to start; all your code is saved for later
- You will be able to run the file you create from within RStudio or
using R’s
source()function.
Tip: Running segments of your code
RStudio offers you great flexibility in running code from within the editor window. There are buttons, menu choices, and keyboard shortcuts. To run the current line, you can
- click on the
Runbutton above the editor panel, or - select “Run Lines” from the “Code” menu, or
- hit Ctrl+Return in Windows or Linux or
⌘+Return on OS X. (This shortcut can also be seen
by hovering the mouse over the button). To run a block of code, select
it and then
Run. If you have modified a line of code within a block of code you have just run, there is no need to reselect the section andRun, you can use the next button along,Re-run the previous region. This will run the previous code block including the modifications you have made.
Introduction to R
Much of your time in R will be spent in the R interactive console.
This is where you will run all of your code, and can be a useful
environment to try out ideas before adding them to an R script file.
This console in RStudio is the same as the one you would get if you
typed in R in your command-line environment.
The first thing you will see in the R interactive session is a bunch of information, followed by a “>” and a blinking cursor. In many ways this is similar to the shell environment you learned about during the shell lessons: it operates on the same idea of a “Read, evaluate, print loop”: you type in commands, R tries to execute them, and then returns a result.
Using R as a calculator
The simplest thing you could do with R is to do arithmetic:
R
1 + 100
OUTPUT
[1] 101And R will print out the answer, with a preceding “[1]”. [1] is the index of the first element of the line being printed in the console. For more information on indexing vectors, see Episode 6: Subsetting Data.
If you type in an incomplete command, R will wait for you to complete it. If you are familiar with Unix Shell’s bash, you may recognize this behavior from bash.
OUTPUT
+Any time you hit return and the R session shows a “+” instead of a “>”, it means it’s waiting for you to complete the command. If you want to cancel a command you can hit Esc and RStudio will give you back the “>” prompt.
Tip: Canceling commands
If you’re using R from the command line instead of from within RStudio, you need to use Ctrl+C instead of Esc to cancel the command. This applies to Mac users as well!
Canceling a command isn’t only useful for killing incomplete commands: you can also use it to tell R to stop running code (for example if it’s taking much longer than you expect), or to get rid of the code you’re currently writing.
When using R as a calculator, the order of operations is the same as you would have learned back in school.
From highest to lowest precedence:
- Parentheses:
(,) - Exponents:
^or** - Multiply:
* - Divide:
/ - Add:
+ - Subtract:
-
R
3 + 5 * 2
OUTPUT
[1] 13Use parentheses to group operations in order to force the order of evaluation if it differs from the default, or to make clear what you intend.
R
(3 + 5) * 2
OUTPUT
[1] 16This can get unwieldy when not needed, but clarifies your intentions. Remember that others may later read your code.
R
(3 + (5 * (2 ^ 2))) # hard to read
3 + 5 * 2 ^ 2 # clear, if you remember the rules
3 + 5 * (2 ^ 2) # if you forget some rules, this might help
The text after each line of code is called a “comment”. Anything that
follows after the hash (or octothorpe) symbol # is ignored
by R when it executes code.
Really small or large numbers get a scientific notation:
R
2/10000
OUTPUT
[1] 2e-04Which is shorthand for “multiplied by 10^XX”. So
2e-4 is shorthand for 2 * 10^(-4).
You can write numbers in scientific notation too:
R
5e3 # Note the lack of minus here
OUTPUT
[1] 5000Mathematical functions
R has many built in mathematical functions. To call a function, we can type its name, followed by open and closing parentheses. Functions take arguments as inputs, anything we type inside the parentheses of a function is considered an argument. Depending on the function, the number of arguments can vary from none to multiple. For example:
R
getwd() #returns an absolute filepath
doesn’t require an argument, whereas for the next set of mathematical functions we will need to supply the function a value in order to compute the result.
R
sin(1) # trigonometry functions
OUTPUT
[1] 0.841471R
log(1) # natural logarithm
OUTPUT
[1] 0R
log10(10) # base-10 logarithm
OUTPUT
[1] 1R
exp(0.5) # e^(1/2)
OUTPUT
[1] 1.648721Don’t worry about trying to remember every function in R. You can look them up on Google, or if you can remember the start of the function’s name, use the tab completion in RStudio.
This is one advantage that RStudio has over R on its own, it has auto-completion abilities that allow you to more easily look up functions, their arguments, and the values that they take.
Typing a ? before the name of a command will open the
help page for that command. When using RStudio, this will open the
‘Help’ pane; if using R in the terminal, the help page will open in your
browser. The help page will include a detailed description of the
command and how it works. Scrolling to the bottom of the help page will
usually show a collection of code examples which illustrate command
usage. We’ll go through an example later.
Comparing things
We can also do comparisons in R:
R
1 == 1 # equality (note two equals signs, read as "is equal to")
OUTPUT
[1] TRUER
1 != 2 # inequality (read as "is not equal to")
OUTPUT
[1] TRUER
1 < 2 # less than
OUTPUT
[1] TRUER
1 <= 1 # less than or equal to
OUTPUT
[1] TRUER
1 > 0 # greater than
OUTPUT
[1] TRUER
1 >= -9 # greater than or equal to
OUTPUT
[1] TRUETip: Comparing Numbers
A word of warning about comparing numbers: you should never use
== to compare two numbers unless they are integers (a data
type which can specifically represent only whole numbers).
Computers may only represent decimal numbers with a certain degree of precision, so two numbers which look the same when printed out by R, may actually have different underlying representations and therefore be different by a small margin of error (called Machine numeric tolerance).
Instead you should use the all.equal function.
Further reading: http://floating-point-gui.de/
Variables and assignment
We can store values in variables using the assignment operator
<-, like this:
R
x <- 1/40
Notice that assignment does not print a value. Instead, we stored it
for later in something called a variable.
x now contains the value
0.025:
R
x
OUTPUT
[1] 0.025More precisely, the stored value is a decimal approximation of this fraction called a floating point number.
Look for the Environment tab in the top right panel of
RStudio, and you will see that x and its value have
appeared. Our variable x can be used in place of a number
in any calculation that expects a number:
R
log(x)
OUTPUT
[1] -3.688879Notice also that variables can be reassigned:
R
x <- 100
x used to contain the value 0.025 and now it has the
value 100.
Assignment values can contain the variable being assigned to:
R
x <- x + 1 #notice how RStudio updates its description of x on the top right tab
y <- x * 2
The right hand side of the assignment can be any valid R expression. The right hand side is fully evaluated before the assignment occurs.
Variable names can contain letters, numbers, underscores and periods but no spaces. They must start with a letter or a period followed by a letter (they cannot start with a number nor an underscore). Variables beginning with a period are hidden variables. Different people use different conventions for long variable names, these include
- periods.between.words
- underscores_between_words
- camelCaseToSeparateWords
What you use is up to you, but be consistent.
It is also possible to use the = operator for
assignment:
R
x = 1/40
But this is much less common among R users. The most important thing
is to be consistent with the operator you use. There
are occasionally places where it is less confusing to use
<- than =, and it is the most common symbol
used in the community. So the recommendation is to use
<-.
Vectorization
One final thing to be aware of is that R is vectorized, meaning that variables and functions can have vectors as values. In contrast to physics and mathematics, a vector in R describes a set of values in a certain order of the same data type. For example:
R
1:5
OUTPUT
[1] 1 2 3 4 5R
2^(1:5)
OUTPUT
[1] 2 4 8 16 32R
x <- 1:5
2^x
OUTPUT
[1] 2 4 8 16 32This is incredibly powerful; we will discuss this further in an upcoming lesson.
Managing your environment
There are a few useful commands you can use to interact with the R session.
ls will list all of the variables and functions stored
in the global environment (your working R session):
R
ls()
OUTPUT
[1] "x" "y"Note here that we didn’t give any arguments to ls, but
we still needed to give the parentheses to tell R to call the
function.
If we type ls by itself, R prints a bunch of code
instead of a listing of objects.
R
ls
OUTPUT
function (name, pos = -1L, envir = as.environment(pos), all.names = FALSE,
pattern, sorted = TRUE)
{
if (!missing(name)) {
pos <- tryCatch(name, error = function(e) e)
if (inherits(pos, "error")) {
name <- substitute(name)
if (!is.character(name))
name <- deparse(name)
warning(gettextf("%s converted to character string",
sQuote(name)), domain = NA)
pos <- name
}
}
all.names <- .Internal(ls(envir, all.names, sorted))
if (!missing(pattern)) {
if ((ll <- length(grep("[", pattern, fixed = TRUE))) &&
ll != length(grep("]", pattern, fixed = TRUE))) {
if (pattern == "[") {
pattern <- "\\["
warning("replaced regular expression pattern '[' by '\\\\['")
}
else if (length(grep("[^\\\\]\\[<-", pattern))) {
pattern <- sub("\\[<-", "\\\\\\[<-", pattern)
warning("replaced '[<-' by '\\\\[<-' in regular expression pattern")
}
}
grep(pattern, all.names, value = TRUE)
}
else all.names
}
<bytecode: 0x55f1cc43f310>
<environment: namespace:base>What’s going on here?
Like everything in R, ls is the name of an object, and
entering the name of an object by itself prints the contents of the
object. The object x that we created earlier contains 1, 2,
3, 4, 5:
R
x
OUTPUT
[1] 1 2 3 4 5The object ls contains the R code that makes the
ls function work! We’ll talk more about how functions work
and start writing our own later.
You can use rm to delete objects you no longer need:
R
rm(x)
If you have lots of things in your environment and want to delete all
of them, you can pass the results of ls to the
rm function:
R
rm(list = ls())
In this case we’ve combined the two. Like the order of operations, anything inside the innermost parentheses is evaluated first, and so on.
In this case we’ve specified that the results of ls
should be used for the list argument in rm.
When assigning values to arguments by name, you must use the
= operator!!
If instead we use <-, there will be unintended side
effects, or you may get an error message:
R
rm(list <- ls())
ERROR
Error in `rm()`:
! ... must contain names or character stringsTip: Warnings vs. Errors
Pay attention when R does something unexpected! Errors, like above, are thrown when R cannot proceed with a calculation. Warnings on the other hand usually mean that the function has run, but it probably hasn’t worked as expected.
In both cases, the message that R prints out usually give you clues how to fix a problem.
R Packages
It is possible to add functions to R by writing a package, or by obtaining a package written by someone else. As of this writing, there are over 10,000 packages available on CRAN (the comprehensive R archive network). R and RStudio have functionality for managing packages:
- You can see what packages are installed by typing
installed.packages() - You can install packages by typing
install.packages("packagename"), wherepackagenameis the package name, in quotes. - You can update installed packages by typing
update.packages() - You can remove a package with
remove.packages("packagename") - You can make a package available for use with
library(packagename)
Packages can also be viewed, loaded, and detached in the Packages tab of the lower right panel in RStudio. Clicking on this tab will display all of the installed packages with a checkbox next to them. If the box next to a package name is checked, the package is loaded and if it is empty, the package is not loaded. Click an empty box to load that package and click a checked box to detach that package.
Packages can be installed and updated from the Package tab with the Install and Update buttons at the top of the tab.
Challenge 2
What will be the value of each variable after each statement in the following program?
R
mass <- 47.5
age <- 122
mass <- mass * 2.3
age <- age - 20
R
mass <- 47.5
This will give a value of 47.5 for the variable mass
R
age <- 122
This will give a value of 122 for the variable age
R
mass <- mass * 2.3
This will multiply the existing value of 47.5 by 2.3 to give a new value of 109.25 to the variable mass.
R
age <- age - 20
This will subtract 20 from the existing value of 122 to give a new value of 102 to the variable age.
Challenge 3
Run the code from the previous challenge, and write a command to compare mass to age. Is mass larger than age?
One way of answering this question in R is to use the
> to set up the following:
R
mass > age
OUTPUT
[1] TRUEThis should yield a boolean value of TRUE since 109.25 is greater than 102.
Challenge 4
Clean up your working environment by deleting the mass and age variables.
We can use the rm command to accomplish this task
R
rm(age, mass)
Challenge 5
Install the following packages: ggplot2,
plyr, gapminder
We can use the install.packages() command to install the
required packages.
R
install.packages("ggplot2")
install.packages("plyr")
install.packages("gapminder")
An alternate solution, to install multiple packages with a single
install.packages() command is:
R
install.packages(c("ggplot2", "plyr", "gapminder"))
When installing ggplot2, it may be required for some users to use the dependencies flag as a result of lazy loading affecting the install. This suggestion is not tied to any known bug discussion, and is advised based off instructor feedback/experience in resolving stochastic occurences of errors identified through delivery of this workshop:
R
install.packages("ggplot2", dependencies = TRUE)
- Use RStudio to write and run R programs.
- R has the usual arithmetic operators and mathematical functions.
- Use
<-to assign values to variables. - Use
ls()to list the variables in a program. - Use
rm()to delete objects in a program. - Use
install.packages()to install packages (libraries).
Content from Project Management With RStudio
Last updated on 2026-04-21 | Edit this page
Estimated time: 30 minutes
Overview
Questions
- How can I manage my projects in R?
Objectives
- Create self-contained projects in RStudio
Introduction
The scientific process is naturally incremental, and many projects start life as random notes, some code, then a manuscript, and eventually everything is a bit mixed together.
Managing your projects in a reproducible fashion doesn’t just make your science reproducible, it makes your life easier.
— Vince Buffalo (@vsbuffalo) April 15, 2013
Most people tend to organize their projects like this:
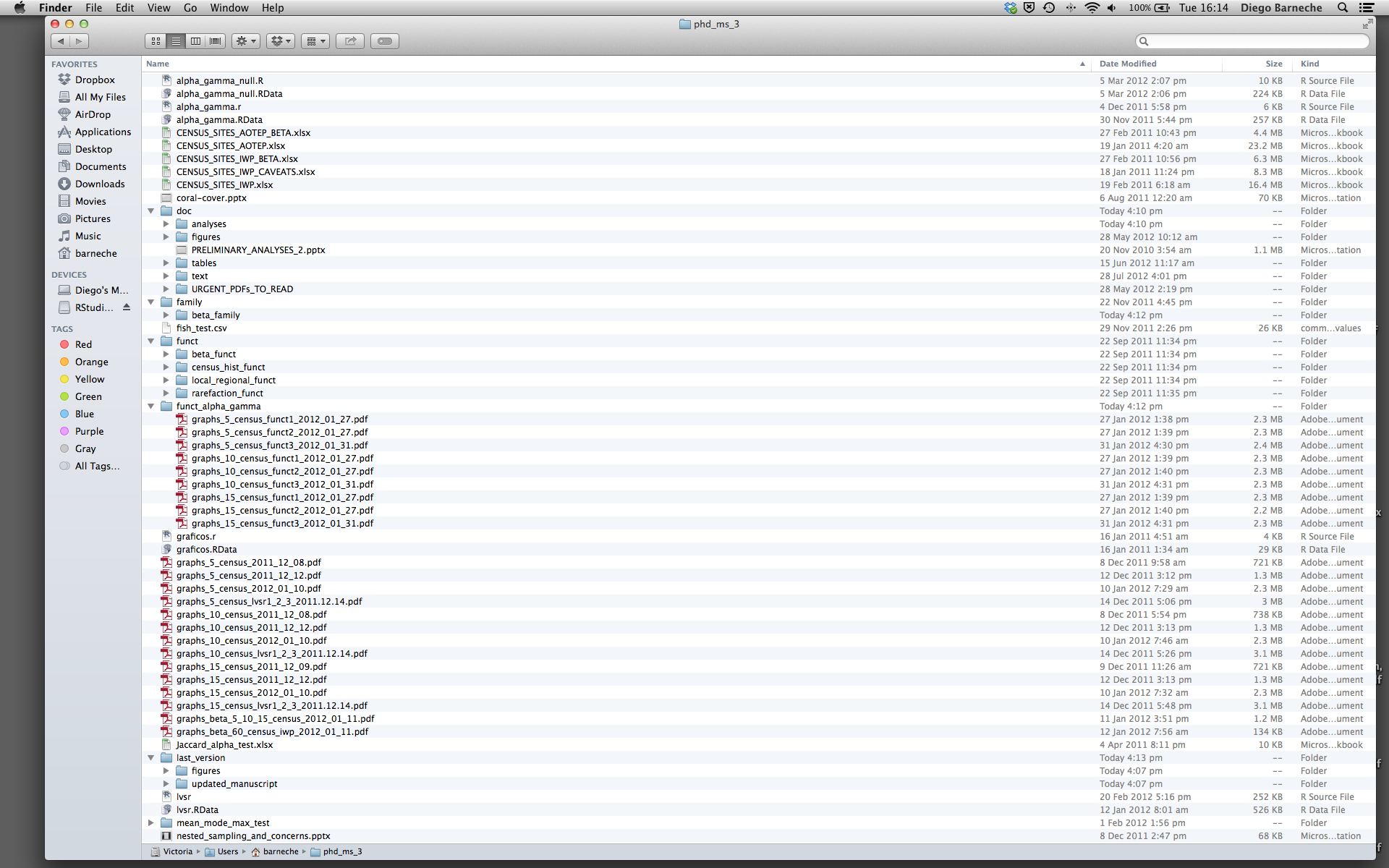
There are many reasons why we should ALWAYS avoid this:
- It is really hard to tell which version of your data is the original and which is the modified;
- It gets really messy because it mixes files with various extensions together;
- It probably takes you a lot of time to actually find things, and relate the correct figures to the exact code that has been used to generate it;
A good project layout will ultimately make your life easier:
- It will help ensure the integrity of your data;
- It makes it simpler to share your code with someone else (a lab-mate, collaborator, or supervisor);
- It allows you to easily upload your code with your manuscript submission;
- It makes it easier to pick the project back up after a break.
A possible solution
Fortunately, there are tools and packages which can help you manage your work effectively.
One of the most powerful and useful aspects of RStudio is its project management functionality. We’ll be using this today to create a self-contained, reproducible project.
Challenge 1: Creating a self-contained project
We’re going to create a new project in RStudio:
- Click the “File” menu button, then “New Project”.
- Click “New Directory”.
- Click “New Project”.
- Type in the name of the directory to store your project, e.g. “my_project”.
- If available, select the checkbox for “Create a git repository.”
- Click the “Create Project” button.
The simplest way to open an RStudio project once it has been created
is to click through your file system to get to the directory where it
was saved and double click on the .Rproj file. This will
open RStudio and start your R session in the same directory as the
.Rproj file. All your data, plots and scripts will now be
relative to the project directory. RStudio projects have the added
benefit of allowing you to open multiple projects at the same time each
open to its own project directory. This allows you to keep multiple
projects open without them interfering with each other.
Challenge 2: Opening an RStudio project through the file system
- Exit RStudio.
- Navigate to the directory where you created a project in Challenge 1.
- Double click on the
.Rprojfile in that directory.
Best practices for project organization
Although there is no “best” way to lay out a project, there are some general principles to adhere to that will make project management easier:
Treat data as read only
This is probably the most important goal of setting up a project. Data is typically time consuming and/or expensive to collect. Working with them interactively (e.g., in Excel) where they can be modified means you are never sure of where the data came from, or how it has been modified since collection. It is therefore a good idea to treat your data as “read-only”.
Data Cleaning
In many cases your data will be “dirty”: it will need significant preprocessing to get into a format R (or any other programming language) will find useful. This task is sometimes called “data munging”. Storing these scripts in a separate folder, and creating a second “read-only” data folder to hold the “cleaned” data sets can prevent confusion between the two sets.
Treat generated output as disposable
Anything generated by your scripts should be treated as disposable: it should all be able to be regenerated from your scripts.
There are lots of different ways to manage this output. Having an output folder with different sub-directories for each separate analysis makes it easier later. Since many analyses are exploratory and don’t end up being used in the final project, and some of the analyses get shared between projects.
Tip: Good Enough Practices for Scientific Computing
Good Enough Practices for Scientific Computing gives the following recommendations for project organization:
- Put each project in its own directory, which is named after the project.
- Put text documents associated with the project in the
docdirectory. - Put raw data and metadata in the
datadirectory, and files generated during cleanup and analysis in aresultsdirectory. - Put source for the project’s scripts and programs in the
srcdirectory, and programs brought in from elsewhere or compiled locally in thebindirectory. - Name all files to reflect their content or function.
Separate function definition and application
One of the more effective ways to work with R is to start by writing the code you want to run directly in a .R script, and then running the selected lines (either using the keyboard shortcuts in RStudio or clicking the “Run” button) in the interactive R console.
When your project is in its early stages, the initial .R script file usually contains many lines of directly executed code. As it matures, reusable chunks get pulled into their own functions. It’s a good idea to separate these functions into two separate folders; one to store useful functions that you’ll reuse across analyses and projects, and one to store the analysis scripts.
Save the data in the data directory
Now we have a good directory structure we will now place/save the
data file in the data/ directory.
Challenge 3
Download the gapminder data from this link to a csv file.
- Download the file (right mouse click on the link above -> “Save link as” / “Save file as”, or click on the link and after the page loads, press Ctrl+S or choose File -> “Save page as”)
- Make sure it’s saved under the name
gapminder_data.csv - Save the file in the
data/folder within your project.
We will load and inspect these data later.
Challenge 4
It is useful to get some general idea about the dataset, directly from the command line, before loading it into R. Understanding the dataset better will come in handy when making decisions on how to load it in R. Use the command-line shell to answer the following questions:
- What is the size of the file?
- How many rows of data does it contain?
- What kinds of values are stored in this file?
By running these commands in the shell:
OUTPUT
-rw-r--r-- 1 root root 80K Apr 21 00:40 data/gapminder_data.csvThe file size is 80K.
OUTPUT
1705 data/gapminder_data.csvThere are 1705 lines. The data looks like:
OUTPUT
country,year,pop,continent,lifeExp,gdpPercap
Afghanistan,1952,8425333,Asia,28.801,779.4453145
Afghanistan,1957,9240934,Asia,30.332,820.8530296
Afghanistan,1962,10267083,Asia,31.997,853.10071
Afghanistan,1967,11537966,Asia,34.02,836.1971382
Afghanistan,1972,13079460,Asia,36.088,739.9811058
Afghanistan,1977,14880372,Asia,38.438,786.11336
Afghanistan,1982,12881816,Asia,39.854,978.0114388
Afghanistan,1987,13867957,Asia,40.822,852.3959448
Afghanistan,1992,16317921,Asia,41.674,649.3413952Tip: command line in RStudio
The Terminal tab in the console pane provides a convenient place directly within RStudio to interact directly with the command line.
Working directory
Knowing R’s current working directory is important because when you need to access other files (for example, to import a data file), R will look for them relative to the current working directory.
Each time you create a new RStudio Project, it will create a new
directory for that project. When you open an existing
.Rproj file, it will open that project and set R’s working
directory to the folder that file is in.
Challenge 5
You can check the current working directory with the
getwd() command, or by using the menus in RStudio.
- In the console, type
getwd()(“wd” is short for “working directory”) and hit Enter. - In the Files pane, double click on the
datafolder to open it (or navigate to any other folder you wish). To get the Files pane back to the current working directory, click “More” and then select “Go To Working Directory”.
You can change the working directory with setwd(), or by
using RStudio menus.
- In the console, type
setwd("data")and hit Enter. Typegetwd()and hit Enter to see the new working directory. - In the menus at the top of the RStudio window, click the “Session”
menu button, and then select “Set Working Directory” and then “Choose
Directory”. Next, in the windows navigator that opens, navigate back to
the project directory, and click “Open”. Note that a
setwdcommand will automatically appear in the console.
Tip: File does not exist errors
When you’re attempting to reference a file in your R code and you’re getting errors saying the file doesn’t exist, it’s a good idea to check your working directory. You need to either provide an absolute path to the file, or you need to make sure the file is saved in the working directory (or a subfolder of the working directory) and provide a relative path.
Version Control
It is important to use version control with projects. Go here for a good lesson which describes using Git with RStudio.
- Use RStudio to create and manage projects with consistent layout.
- Treat raw data as read-only.
- Treat generated output as disposable.
- Separate function definition and application.
Content from Seeking Help
Last updated on 2026-04-21 | Edit this page
Estimated time: 20 minutes
Overview
Questions
- How can I get help in R?
Objectives
- To be able to read R help files for functions and special operators.
- To be able to use CRAN task views to identify packages to solve a problem.
- To be able to seek help from your peers.
Reading Help Files
R, and every package, provide help files for functions. The general syntax to search for help on any function, “function_name”, from a specific function that is in a package loaded into your namespace (your interactive R session) is:
R
?function_name
help(function_name)
For example take a look at the help file for
write.table(), we will be using a similar function in an
upcoming episode.
R
?write.table()
This will load up a help page in RStudio (or as plain text in R itself).
Each help page is broken down into sections:
- Description: An extended description of what the function does.
- Usage: The arguments of the function and their default values (which can be changed).
- Arguments: An explanation of the data each argument is expecting.
- Details: Any important details to be aware of.
- Value: The data the function returns.
- See Also: Any related functions you might find useful.
- Examples: Some examples for how to use the function.
Different functions might have different sections, but these are the main ones you should be aware of.
Notice how related functions might call for the same help file:
R
?write.table()
?write.csv()
This is because these functions have very similar applicability and often share the same arguments as inputs to the function, so package authors often choose to document them together in a single help file.
Tip: Running Examples
From within the function help page, you can highlight code in the Examples and hit Ctrl+Return to run it in RStudio console. This gives you a quick way to get a feel for how a function works.
Tip: Reading Help Files
One of the most daunting aspects of R is the large number of functions available. It would be prohibitive, if not impossible to remember the correct usage for every function you use. Luckily, using the help files means you don’t have to remember that!
Special Operators
To seek help on special operators, use quotes or backticks:
R
?"<-"
?`<-`
Getting Help with Packages
Many packages come with “vignettes”: tutorials and extended example
documentation. Without any arguments, vignette() will list
all vignettes for all installed packages;
vignette(package="package-name") will list all available
vignettes for package-name, and
vignette("vignette-name") will open the specified
vignette.
If a package doesn’t have any vignettes, you can usually find help by
typing help("package-name").
RStudio also has a set of excellent cheatsheets for many packages.
When You Remember Part of the Function Name
If you’re not sure what package a function is in or how it’s specifically spelled, you can do a fuzzy search:
R
??function_name
A fuzzy search is when you search for an approximate string match. For example, you may remember that the function to set your working directory includes “set” in its name. You can do a fuzzy search to help you identify the function:
R
??set
When You Have No Idea Where to Begin
If you don’t know what function or package you need to use CRAN Task Views is a specially maintained list of packages grouped into fields. This can be a good starting point.
Other ways to get help
There are several other ways that people often get help when they are stuck with their R code.
- Search the internet: paste the last line of your error message or
“R” and a short description of what you want to do into your favourite
search engine and you will usually find several examples where other
people have encountered the same problem and came looking for help.
- StackOverflow can be particularly helpful for this: answers to
questions are presented as a ranked thread ordered according to how
useful other users found them to be. You can search using the
[r]tag. - Take care: copying and pasting code written by somebody else is risky unless you understand exactly what it is doing!
- StackOverflow can be particularly helpful for this: answers to
questions are presented as a ranked thread ordered according to how
useful other users found them to be. You can search using the
- Ask somebody “in the real world”. If you have a colleague or friend with more expertise in R than you have, show them the problem you are having and ask them for help.
- Sometimes, the act of articulating your question can help you to identify what is going wrong. This is known as “rubber duck debugging” among programmers.
Formulating your question
The StackOverflow page on how to ask a good question has tips on what information to include when posting a question on StackOverflow. Some of these are specific to StackOverflow but others are also relevant if you’re asking a colleague or friend for help, or even if just articulating your question to yourself.
R has a few useful functions to help you formulate your question:
R
?dput
Will dump the data you’re working with into a format that can be copied and pasted by others into their own R session.
R
sessionInfo()
OUTPUT
R version 4.5.3 (2026-03-11)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
time zone: Etc/UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
loaded via a namespace (and not attached):
[1] compiler_4.5.3 cli_3.6.5 tools_4.5.3 otel_0.2.0 yaml_2.3.12
[6] knitr_1.51 xfun_0.57 rlang_1.2.0 renv_1.2.2 evaluate_1.0.5Will print out your current version of R, as well as any packages you have loaded. This can be useful for others to help reproduce and debug your issue.
Generative AI
The section on generative AI is intended to be concise but Instructors may choose to devote more time to the topic in a workshop.
Depending on your own level of experience and comfort with talking about and using these tools, you could choose to do any of the following:
Explain how large language models work and are trained, and/or the difference between generative AI, other forms of AI that currently exist, and the limits of what LLMs can do (e.g., they can’t “reason”).
Demonstrate how you recommend that learners use generative AI.
Discuss the ethical concerns listed below, as well as others that you are aware of, to help learners make an informed choice about whether or not to use generative AI tools.
This is a fast-moving technology. If you are preparing to teach this section and you feel it has become outdated, please open an issue on the lesson repository to let the Maintainers know and/or a pull request to suggest updates and improvements.
It is increasingly common for people to use generative AI chatbots such as ChatGPT to get help while coding. You will probably receive some useful guidance by presenting your error message to the chatbot and asking it what went wrong. However, the way this help is provided by the chatbot is different. Answers on StackOverflow have (probably) been given by a human as a direct response to the question asked. But generative AI chatbots, which are based on an advanced statistical model, respond by generating the most likely sequence of text that would follow the prompt they are given.
While responses from generative AI tools can often be helpful, they are not always reliable. These tools sometimes generate plausible but incorrect or misleading information, so (just as with an answer found on the internet) it is essential to verify their accuracy. You need the knowledge and skills to be able to understand these responses, to judge whether or not they are accurate, and to fix any errors in the code it offers you.
In addition to asking for help, programmers can use generative AI tools to generate code from scratch; extend, improve and reorganise existing code; translate code between programming languages; figure out what terms to use in a search of the internet; and more. However, there are drawbacks that you should be aware of.
The models used by these tools have been “trained” on very large volumes of data, much of it taken from the internet, and the responses they produce reflect that training data, and may recapitulate its inaccuracies or biases. The environmental costs (energy and water use) of LLMs are a lot higher than other technologies, both during development (known as training) and when an individual user uses one (also called inference). For more information see the AI Environmental Impact Primer developed by researchers at HuggingFace, an AI hosting platform. Concerns also exist about the way the data for this training was obtained, with questions raised about whether the people developing the LLMs had permission to use it. Other ethical concerns have also been raised, such as reports that workers were exploited during the training process.
We recommend that you avoid getting help from generative AI during the workshop for several reasons:
- For most problems you will encounter at this stage, help and answers can be found among the first results returned by searching the internet.
- The foundational knowledge and skills you will learn in this lesson by writing and fixing your own programs are essential to be able to evaluate the correctness and safety of any code you receive from online help or a generative AI chatbot. If you choose to use these tools in the future, the expertise you gain from learning and practising these fundamentals on your own will help you use them more effectively.
- As you start out with programming, the mistakes you make will be the kinds that have also been made – and overcome! – by everybody else who learned to program before you. Since these mistakes and the questions you are likely to have at this stage are common, they are also better represented than other, more specialised problems and tasks in the data that was used to train generative AI tools. This means that a generative AI chatbot is more likely to produce accurate responses to questions that novices ask, which could give you a false impression of how reliable they will be when you are ready to do things that are more advanced.
Challenge 1
Look at the help page for the c function. What kind of
vector do you expect will be created if you evaluate the following:
R
c(1, 2, 3)
c('d', 'e', 'f')
c(1, 2, 'f')
The c() function creates a vector, in which all elements
are of the same type. In the first case, the elements are numeric, in
the second, they are characters, and in the third they are also
characters: the numeric values are “coerced” to be characters.
Challenge 2
Look at the help for the paste function. You will need
to use it later. What’s the difference between the sep and
collapse arguments?
To look at the help for the paste() function, use:
R
help("paste")
?paste
The difference between sep and collapse is
a little tricky. The paste function accepts any number of
arguments, each of which can be a vector of any length. The
sep argument specifies the string used between concatenated
terms — by default, a space. The result is a vector as long as the
longest argument supplied to paste. In contrast,
collapse specifies that after concatenation the elements
are collapsed together using the given separator, the result
being a single string.
It is important to call the arguments explicitly by typing out the
argument name e.g sep = "," so the function understands to
use the “,” as a separator and not a term to concatenate. e.g.
R
paste(c("a","b"), "c")
OUTPUT
[1] "a c" "b c"R
paste(c("a","b"), "c", ",")
OUTPUT
[1] "a c ," "b c ,"R
paste(c("a","b"), "c", sep = ",")
OUTPUT
[1] "a,c" "b,c"R
paste(c("a","b"), "c", collapse = "|")
OUTPUT
[1] "a c|b c"R
paste(c("a","b"), "c", sep = ",", collapse = "|")
OUTPUT
[1] "a,c|b,c"(For more information, scroll to the bottom of the
?paste help page and look at the examples, or try
example('paste').)
Challenge 3
Use help to find a function (and its associated parameters) that you
could use to load data from a tabular file in which columns are
delimited with “\t” (tab) and the decimal point is a “.” (period). This
check for decimal separator is important, especially if you are working
with international colleagues, because different countries have
different conventions for the decimal point (i.e. comma vs period).
Hint: use ??"read table" to look up functions related to
reading in tabular data.
The standard R function for reading tab-delimited files with a period
decimal separator is read.delim(). You can also do this with
read.table(file, sep="\t") (the period is the
default decimal separator for read.table()),
although you may have to change the comment.char argument
as well if your data file contains hash (#) characters.
Resources
- Use
help()to get online help in R.
Content from Data Structures
Last updated on 2026-04-21 | Edit this page
Estimated time: 55 minutes
Overview
Questions
- How can I read data in R?
- What are the basic data types in R?
- How do I represent categorical information in R?
Objectives
- To be able to identify the 5 main data types.
- To begin exploring data frames, and understand how they are related to vectors and lists.
- To be able to ask questions from R about the type, class, and structure of an object.
- To understand the information of the attributes “names”, “class”, and “dim”.
One of R’s most powerful features is its ability to deal with tabular
data - such as you may already have in a spreadsheet or a CSV file.
Let’s start by making a toy dataset in your data/
directory, called feline-data.csv:
R
cats <- data.frame(coat = c("calico", "black", "tabby"),
weight = c(2.1, 5.0, 3.2),
likes_catnip = c(1, 0, 1))
We can now save cats as a CSV file. It is good practice
to call the argument names explicitly so the function knows what default
values you are changing. Here we are setting
row.names = FALSE. Recall you can use
?write.csv to pull up the help file to check out the
argument names and their default values.
R
write.csv(x = cats, file = "data/feline-data.csv", row.names = FALSE)
The contents of the new file, feline-data.csv:
Tip: Editing Text files in R
Alternatively, you can create data/feline-data.csv using
a text editor (Nano), or within RStudio with the File -> New
File -> Text File menu item.
We can load this into R via the following:
R
cats <- read.csv(file = "data/feline-data.csv")
cats
OUTPUT
coat weight likes_catnip
1 calico 2.1 1
2 black 5.0 0
3 tabby 3.2 1The read.csv function is used for reading in tabular
data stored in a text file where the columns of data are separated by
punctuation characters such as CSV files (csv = comma-separated values).
Tabs and commas are the most common punctuation characters used to
separate or delimit data points in csv files. For convenience R provides
2 other versions of read.table. These are:
read.csv for files where the data are separated with commas
and read.delim for files where the data are separated with
tabs. Of these three functions read.csv is the most
commonly used. If needed it is possible to override the default
delimiting punctuation marks for both read.csv and
read.delim.
Check your data for factors
In recent times, the default way how R handles textual data has changed. Text data was interpreted by R automatically into a format called “factors”. But there is an easier format that is called “character”. We will hear about factors later, and what to use them for. For now, remember that in most cases, they are not needed and only complicate your life, which is why newer R versions read in text as “character”. Check now if your version of R has automatically created factors and convert them to “character” format:
- Check the data types of your input by typing
str(cats) - In the output, look at the three-letter codes after the colons: If you see only “num” and “chr”, you can continue with the lesson and skip this box. If you find “fct”, continue to step 3.
- Prevent R from automatically creating “factor” data. That can be
done by the following code:
options(stringsAsFactors = FALSE). Then, re-read the cats table for the change to take effect. - You must set this option every time you restart R. To not forget this, include it in your analysis script before you read in any data, for example in one of the first lines.
- For R versions greater than 4.0.0, text data is no longer converted to factors anymore. So you can install this or a newer version to avoid this problem. If you are working on an institute or company computer, ask your administrator to do it.
We can begin exploring our dataset right away, pulling out columns by
specifying them using the $ operator:
R
cats$weight
OUTPUT
[1] 2.1 5.0 3.2R
cats$coat
OUTPUT
[1] "calico" "black" "tabby" We can do other operations on the columns:
R
## Say we discovered that the scale weighs two Kg light:
cats$weight + 2
OUTPUT
[1] 4.1 7.0 5.2R
paste("My cat is", cats$coat)
OUTPUT
[1] "My cat is calico" "My cat is black" "My cat is tabby" But what about
R
cats$weight + cats$coat
ERROR
Error in `cats$weight + cats$coat`:
! non-numeric argument to binary operatorUnderstanding what happened here is key to successfully analyzing data in R.
Data Types
If you guessed that the last command will return an error because
2.1 plus "black" is nonsense, you’re right -
and you already have some intuition for an important concept in
programming called data types. We can ask what type of data
something is:
R
typeof(cats$weight)
OUTPUT
[1] "double"There are 5 main types: double, integer,
complex, logical and character.
For historic reasons, double is also called
numeric.
R
typeof(3.14)
OUTPUT
[1] "double"R
typeof(1L) # The L suffix forces the number to be an integer, since by default R uses float numbers
OUTPUT
[1] "integer"R
typeof(1+1i)
OUTPUT
[1] "complex"R
typeof(TRUE)
OUTPUT
[1] "logical"R
typeof('banana')
OUTPUT
[1] "character"No matter how complicated our analyses become, all data in R is interpreted as one of these basic data types. This strictness has some really important consequences.
A user has provided details of another cat. We can add an additional
row to our cats table using rbind().
R
additional_cat <- data.frame(coat = "tabby", weight = "2.3 or 2.4", likes_catnip = 1)
additional_cat
OUTPUT
coat weight likes_catnip
1 tabby 2.3 or 2.4 1R
cats2 <- rbind(cats, additional_cat)
cats2
OUTPUT
coat weight likes_catnip
1 calico 2.1 1
2 black 5 0
3 tabby 3.2 1
4 tabby 2.3 or 2.4 1Let’s check what type of data we find in the weight
column of our new cats2 object:
R
typeof(cats2$weight)
OUTPUT
[1] "character"Oh no, our weights aren’t the double type anymore! If we try to do the same math we did on them before, we run into trouble:
R
cats2$weight + 2
ERROR
Error in `cats2$weight + 2`:
! non-numeric argument to binary operatorWhat happened? The cats (and cats2) data we
are working with is something called a data frame. Data frames
are one of the most common and versatile types of data
structures we will work with in R. A given column in a data frame
cannot be composed of different data types. In this case, R cannot store
everything in the data frame column weight as a
double anymore once we add the row for the additional cat
(because its weight is 2.3 or 2.4), therefore the entire
column data type changes to something that is suitable for everything in
the column.
When R reads a csv file, it reads it in as a data frame.
Thus, when we loaded the cats csv file, it is stored as a
data frame. We can recognize data frames by the first row that is
written by the str() function:
R
str(cats2)
OUTPUT
'data.frame': 4 obs. of 3 variables:
$ coat : chr "calico" "black" "tabby" "tabby"
$ weight : chr "2.1" "5" "3.2" "2.3 or 2.4"
$ likes_catnip: num 1 0 1 1Data frames are composed of rows and columns, where each column has the same number of rows. Different columns in a data frame can be made up of different data types (this is what makes them so versatile), but everything in a given column needs to be the same type (e.g., vector, factor, or list).
Let’s explore more about different data structures and how they
behave. For now, we will focus on our original data frame
cats (and we can forget about cats2 for the
rest of this episode).
Vectors and Type Coercion
To better understand this behavior, let’s meet another of the data structures: the vector.
R
my_vector <- vector(length = 3)
my_vector
OUTPUT
[1] FALSE FALSE FALSEA vector in R is essentially an ordered list of things, with the
special condition that everything in the vector must be the same
basic data type. If you don’t choose the datatype, it’ll default to
logical; or, you can declare an empty vector of whatever
type you like.
R
another_vector <- vector(mode='character', length=3)
another_vector
OUTPUT
[1] "" "" ""You can check if something is a vector:
R
str(another_vector)
OUTPUT
chr [1:3] "" "" ""The somewhat cryptic output from this command indicates the basic
data type found in this vector - in this case chr,
character; an indication of the number of things in the vector -
actually, the indexes of the vector, in this case [1:3];
and a few examples of what’s actually in the vector - in this case empty
character strings. If we similarly do
R
str(cats$weight)
OUTPUT
num [1:3] 2.1 5 3.2we see that cats$weight is a vector, too - the
columns of data we load into R data.frames are all vectors, and
that’s the root of why R forces everything in a column to be the same
basic data type.
Discussion 1
Why is R so opinionated about what we put in our columns of data? How does this help us?
By keeping everything in a column the same, we allow ourselves to make simple assumptions about our data; if you can interpret one entry in the column as a number, then you can interpret all of them as numbers, so we don’t have to check every time. This consistency is what people mean when they talk about clean data; in the long run, strict consistency goes a long way to making our lives easier in R.
Coercion by combining vectors
You can also make vectors with explicit contents with the combine function:
R
combine_vector <- c(2,6,3)
combine_vector
OUTPUT
[1] 2 6 3Given what we’ve learned so far, what do you think the following will produce?
R
quiz_vector <- c(2,6,'3')
This is something called type coercion, and it is the source of many surprises and the reason why we need to be aware of the basic data types and how R will interpret them. When R encounters a mix of types (here double and character) to be combined into a single vector, it will force them all to be the same type. Consider:
R
coercion_vector <- c('a', TRUE)
coercion_vector
OUTPUT
[1] "a" "TRUE"R
another_coercion_vector <- c(0, TRUE)
another_coercion_vector
OUTPUT
[1] 0 1The type hierarchy
The coercion rules go: logical ->
integer -> double (“numeric”)
-> complex -> character, where -> can
be read as are transformed into. For example, combining
logical and character transforms the result to
character:
R
c('a', TRUE)
OUTPUT
[1] "a" "TRUE"A quick way to recognize character vectors is by the
quotes that enclose them when they are printed.
You can try to force coercion against this flow using the
as. functions:
R
character_vector_example <- c('0','2','4')
character_vector_example
OUTPUT
[1] "0" "2" "4"R
character_coerced_to_double <- as.double(character_vector_example)
character_coerced_to_double
OUTPUT
[1] 0 2 4R
double_coerced_to_logical <- as.logical(character_coerced_to_double)
double_coerced_to_logical
OUTPUT
[1] FALSE TRUE TRUEAs you can see, some surprising things can happen when R forces one basic data type into another! Nitty-gritty of type coercion aside, the point is: if your data doesn’t look like what you thought it was going to look like, type coercion may well be to blame; make sure everything is the same type in your vectors and your columns of data.frames, or you will get nasty surprises!
But coercion can also be very useful! For example, in our
cats data likes_catnip is numeric, but we know
that the 1s and 0s actually represent TRUE and
FALSE (a common way of representing them). We should use
the logical datatype here, which has two states:
TRUE or FALSE, which is exactly what our data
represents. We can ‘coerce’ this column to be logical by
using the as.logical function:
R
cats$likes_catnip
OUTPUT
[1] 1 0 1R
cats$likes_catnip <- as.logical(cats$likes_catnip)
cats$likes_catnip
OUTPUT
[1] TRUE FALSE TRUEChallenge 1
An important part of every data analysis is cleaning the input data. If you know that the input data is all of the same format, (e.g. numbers), your analysis is much easier! Clean the cat data set from the chapter about type coercion.
Copy the code template
Create a new script in RStudio and copy and paste the following code. Then move on to the tasks below, which help you to fill in the gaps (______).
Using the object `cats2`:
# 1. Print the data
_____
# 2. Show an overview of the table with all data types
_____(cats2)
# 3. The "weight" column has the incorrect data type __________.
# The correct data type is: ____________.
# 4. Correct the 4th weight data point with the mean of the two given values
cats2$weight[4] <- 2.35
# print the data again to see the effect
cats2
# 5. Convert the weight to the right data type
cats2$weight <- ______________(cats2$weight)
# Calculate the mean to test yourself
mean(cats2$weight)
# If you see the correct mean value (and not NA), you did the exercise
# correctly!2. Overview of the data types
The data type of your data is as important as the data itself. Use a
function we saw earlier to print out the data types of all columns of
the cats2 data.frame.
In the chapter “Data types” we saw two functions that can show data types. One printed just a single word, the data type name. The other printed a short form of the data type, and the first few values. We need the second here.
Scroll up to the section about the type hierarchy to review the available data types
- Weight is expressed on a continuous scale (real numbers). The R data type for this is “double” (also known as “numeric”).
- The fourth row has the value “2.3 or 2.4”. That is not a number but two, and an english word. Therefore, the “character” data type is chosen. The whole column is now text, because all values in the same columns have to be the same data type.
4. Correct the problematic value
The code to assign a new weight value to the problematic fourth row is given. Think first and then execute it: What will be the data type after assigning a number like in this example? You can check the data type after executing to see if you were right.
Revisit the hierarchy of data types when two different data types are combined.
Challenge 1 (continued)
Solution to challenge 1.4
The data type of the column “weight” is “character”. The assigned data type is “double”. Combining two data types yields the data type that is higher in the following hierarchy:
logical < integer < double < complex < characterTherefore, the column is still of type character! We need to manually convert it to “double”. {: .solution}
The functions to convert data types start with as.. You
can look for the function further up in the manuscript or use the
RStudio auto-complete function: Type “as.” and then press
the TAB key.
Challenge 1 (continued)
Solution to Challenge 1.5
There are two functions that are synonymous for historic reasons:
cats2$weight <- as.double(cats2$weight) cats2$weight <- as.numeric(cats2$weight)
Some basic vector functions
The combine function, c(), will also append things to an
existing vector:
R
ab_vector <- c('a', 'b')
ab_vector
OUTPUT
[1] "a" "b"R
combine_example <- c(ab_vector, 'SWC')
combine_example
OUTPUT
[1] "a" "b" "SWC"You can also make series of numbers:
R
mySeries <- 1:10
mySeries
OUTPUT
[1] 1 2 3 4 5 6 7 8 9 10R
seq(10)
OUTPUT
[1] 1 2 3 4 5 6 7 8 9 10R
seq(1,10, by=0.1)
OUTPUT
[1] 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 1.8 1.9 2.0 2.1 2.2 2.3 2.4
[16] 2.5 2.6 2.7 2.8 2.9 3.0 3.1 3.2 3.3 3.4 3.5 3.6 3.7 3.8 3.9
[31] 4.0 4.1 4.2 4.3 4.4 4.5 4.6 4.7 4.8 4.9 5.0 5.1 5.2 5.3 5.4
[46] 5.5 5.6 5.7 5.8 5.9 6.0 6.1 6.2 6.3 6.4 6.5 6.6 6.7 6.8 6.9
[61] 7.0 7.1 7.2 7.3 7.4 7.5 7.6 7.7 7.8 7.9 8.0 8.1 8.2 8.3 8.4
[76] 8.5 8.6 8.7 8.8 8.9 9.0 9.1 9.2 9.3 9.4 9.5 9.6 9.7 9.8 9.9
[91] 10.0We can ask a few questions about vectors:
R
sequence_example <- 20:25
head(sequence_example, n=2)
OUTPUT
[1] 20 21R
tail(sequence_example, n=4)
OUTPUT
[1] 22 23 24 25R
length(sequence_example)
OUTPUT
[1] 6R
typeof(sequence_example)
OUTPUT
[1] "integer"We can get individual elements of a vector by using the bracket notation:
R
first_element <- sequence_example[1]
first_element
OUTPUT
[1] 20To change a single element, use the bracket on the other side of the arrow:
R
sequence_example[1] <- 30
sequence_example
OUTPUT
[1] 30 21 22 23 24 25Challenge 2
Start by making a vector with the numbers 1 through 26. Then, multiply the vector by 2.
R
x <- 1:26
x <- x * 2
Lists
Another data structure you’ll want in your bag of tricks is the
list. A list is simpler in some ways than the other types,
because you can put anything you want in it. Remember everything in
the vector must be of the same basic data type, but a list can have
different data types:
R
list_example <- list(1, "a", TRUE, 1+4i)
list_example
OUTPUT
[[1]]
[1] 1
[[2]]
[1] "a"
[[3]]
[1] TRUE
[[4]]
[1] 1+4iWhen printing the object structure with str(), we see
the data types of all elements:
R
str(list_example)
OUTPUT
List of 4
$ : num 1
$ : chr "a"
$ : logi TRUE
$ : cplx 1+4iWhat is the use of lists? They can organize data of different types. For example, you can organize different tables that belong together, similar to spreadsheets in Excel. But there are many other uses, too.
We will see another example that will maybe surprise you in the next chapter.
To retrieve one of the elements of a list, use the double bracket:
R
list_example[[2]]
OUTPUT
[1] "a"The elements of lists also can have names, they can be given by prepending them to the values, separated by an equals sign:
R
another_list <- list(title = "Numbers", numbers = 1:10, data = TRUE )
another_list
OUTPUT
$title
[1] "Numbers"
$numbers
[1] 1 2 3 4 5 6 7 8 9 10
$data
[1] TRUEThis results in a named list. Now we have a new function of our object! We can access single elements by an additional way!
R
another_list$title
OUTPUT
[1] "Numbers"Names
With names, we can give meaning to elements. It is the first time that we do not only have the data, but also explaining information. It is metadata that can be stuck to the object like a label. In R, this is called an attribute. Some attributes enable us to do more with our object, for example, like here, accessing an element by a self-defined name.
Accessing vectors and lists by name
We have already seen how to generate a named list. The way to generate a named vector is very similar. You have seen this function before:
R
pizza_price <- c( pizzasubito = 5.64, pizzafresh = 6.60, callapizza = 4.50 )
The way to retrieve elements is different, though:
R
pizza_price["pizzasubito"]
OUTPUT
pizzasubito
5.64 The approach used for the list does not work:
R
pizza_price$pizzafresh
ERROR
Error in `pizza_price$pizzafresh`:
! $ operator is invalid for atomic vectorsIt will pay off if you remember this error message, you will meet it in your own analyses. It means that you have just tried accessing an element like it was in a list, but it is actually in a vector.
Accessing and changing names
If you are only interested in the names, use the names()
function:
R
names(pizza_price)
OUTPUT
[1] "pizzasubito" "pizzafresh" "callapizza" We have seen how to access and change single elements of a vector. The same is possible for names:
R
names(pizza_price)[3]
OUTPUT
[1] "callapizza"R
names(pizza_price)[3] <- "call-a-pizza"
pizza_price
OUTPUT
pizzasubito pizzafresh call-a-pizza
5.64 6.60 4.50 Challenge 3
- What is the data type of the names of
pizza_price? You can find out using thestr()ortypeof()functions.
You get the names of an object by wrapping the object name inside
names(...). Similarly, you get the data type of the names
by again wrapping the whole code in typeof(...):
typeof(names(pizza))alternatively, use a new variable if this is easier for you to read:
n <- names(pizza)
typeof(n)Challenge 4
Instead of just changing some of the names a vector/list already has, you can also set all names of an object by writing code like (replace ALL CAPS text):
names( OBJECT ) <- CHARACTER_VECTORCreate a vector that gives the number for each letter in the alphabet!
- Generate a vector called
letter_nowith the sequence of numbers from 1 to 26! - R has a built-in object called
LETTERS. It is a 26-character vector, from A to Z. Set the names of the number sequence to this 26 letters - Test yourself by calling
letter_no["B"], which should give you the number 2!
letter_no <- 1:26 # or seq(1,26)
names(letter_no) <- LETTERS
letter_no["B"]Data frames
We have data frames at the very beginning of this lesson, they represent a table of data. We didn’t go much further into detail with our example cat data frame:
R
cats
OUTPUT
coat weight likes_catnip
1 calico 2.1 TRUE
2 black 5.0 FALSE
3 tabby 3.2 TRUEWe can now understand something a bit surprising in our data.frame; what happens if we run:
R
typeof(cats)
OUTPUT
[1] "list"We see that data.frames look like lists ‘under the hood’. Think again what we heard about what lists can be used for:
Lists organize data of different types
Columns of a data frame are vectors of different types, that are organized by belonging to the same table.
A data.frame is really a list of vectors. It is a special list in which all the vectors must have the same length.
How is this “special”-ness written into the object, so that R does not treat it like any other list, but as a table?
R
class(cats)
OUTPUT
[1] "data.frame"A class, just like names, is an attribute attached to the object. It tells us what this object means for humans.
You might wonder: Why do we need another
what-type-of-object-is-this-function? We already have
typeof()? That function tells us how the object is
constructed in the computer. The class is
the meaning of the object for humans. Consequently,
what typeof() returns is fixed in R (mainly the
five data types), whereas the output of class() is
diverse and extendable by R packages.
In our cats example, we have an integer, a double and a
logical variable. As we have seen already, each column of data.frame is
a vector.
R
cats$coat
OUTPUT
[1] "calico" "black" "tabby" R
cats[,1]
OUTPUT
[1] "calico" "black" "tabby" R
typeof(cats[,1])
OUTPUT
[1] "character"R
str(cats[,1])
OUTPUT
chr [1:3] "calico" "black" "tabby"Each row is an observation of different variables, itself a data.frame, and thus can be composed of elements of different types.
R
cats[1,]
OUTPUT
coat weight likes_catnip
1 calico 2.1 TRUER
typeof(cats[1,])
OUTPUT
[1] "list"R
str(cats[1,])
OUTPUT
'data.frame': 1 obs. of 3 variables:
$ coat : chr "calico"
$ weight : num 2.1
$ likes_catnip: logi TRUEChallenge 5
There are several subtly different ways to call variables, observations and elements from data.frames:
cats[1]cats[[1]]cats$coatcats["coat"]cats[1, 1]cats[, 1]cats[1, ]
Try out these examples and explain what is returned by each one.
Hint: Use the function typeof() to examine what
is returned in each case.
R
cats[1]
OUTPUT
coat
1 calico
2 black
3 tabbyWe can think of a data frame as a list of vectors. The single brace
[1] returns the first slice of the list, as another list.
In this case it is the first column of the data frame.
R
cats[[1]]
OUTPUT
[1] "calico" "black" "tabby" The double brace [[1]] returns the contents of the list
item. In this case it is the contents of the first column, a
vector of type character.
R
cats$coat
OUTPUT
[1] "calico" "black" "tabby" This example uses the $ character to address items by
name. coat is the first column of the data frame, again a
vector of type character.
R
cats["coat"]
OUTPUT
coat
1 calico
2 black
3 tabbyHere we are using a single brace ["coat"] replacing the
index number with the column name. Like example 1, the returned object
is a list.
R
cats[1, 1]
OUTPUT
[1] "calico"This example uses a single brace, but this time we provide row and column coordinates. The returned object is the value in row 1, column 1. The object is a vector of type character.
R
cats[, 1]
OUTPUT
[1] "calico" "black" "tabby" Like the previous example we use single braces and provide row and column coordinates. The row coordinate is not specified, R interprets this missing value as all the elements in this column and returns them as a vector.
R
cats[1, ]
OUTPUT
coat weight likes_catnip
1 calico 2.1 TRUEAgain we use the single brace with row and column coordinates. The column coordinate is not specified. The return value is a list containing all the values in the first row.
Tip: Renaming data frame columns
Data frames have column names, which can be accessed with the
names() function.
R
names(cats)
OUTPUT
[1] "coat" "weight" "likes_catnip"If you want to rename the second column of cats, you can
assign a new name to the second element of names(cats).
R
names(cats)[2] <- "weight_kg"
cats
OUTPUT
coat weight_kg likes_catnip
1 calico 2.1 TRUE
2 black 5.0 FALSE
3 tabby 3.2 TRUEMatrices
Last but not least is the matrix. We can declare a matrix full of zeros:
R
matrix_example <- matrix(0, ncol=6, nrow=3)
matrix_example
OUTPUT
[,1] [,2] [,3] [,4] [,5] [,6]
[1,] 0 0 0 0 0 0
[2,] 0 0 0 0 0 0
[3,] 0 0 0 0 0 0What makes it special is the dim() attribute:
R
dim(matrix_example)
OUTPUT
[1] 3 6And similar to other data structures, we can ask things about our matrix:
R
typeof(matrix_example)
OUTPUT
[1] "double"R
class(matrix_example)
OUTPUT
[1] "matrix" "array" R
str(matrix_example)
OUTPUT
num [1:3, 1:6] 0 0 0 0 0 0 0 0 0 0 ...R
nrow(matrix_example)
OUTPUT
[1] 3R
ncol(matrix_example)
OUTPUT
[1] 6Challenge 6
What do you think will be the result of
length(matrix_example)? Try it. Were you right? Why / why
not?
What do you think will be the result of
length(matrix_example)?
R
matrix_example <- matrix(0, ncol=6, nrow=3)
length(matrix_example)
OUTPUT
[1] 18Because a matrix is a vector with added dimension attributes,
length gives you the total number of elements in the
matrix.
Challenge 7
Make another matrix, this time containing the numbers 1:50, with 5
columns and 10 rows. Did the matrix function fill your
matrix by column, or by row, as its default behaviour? See if you can
figure out how to change this. (hint: read the documentation for
matrix!)
Make another matrix, this time containing the numbers 1:50, with 5
columns and 10 rows. Did the matrix function fill your
matrix by column, or by row, as its default behaviour? See if you can
figure out how to change this. (hint: read the documentation for
matrix!)
R
x <- matrix(1:50, ncol=5, nrow=10)
x <- matrix(1:50, ncol=5, nrow=10, byrow = TRUE) # to fill by row
Challenge 8
Create a list of length two containing a character vector for each of the sections in this part of the workshop:
- Data types
- Data structures
Populate each character vector with the names of the data types and data structures we’ve seen so far.
R
dataTypes <- c('double', 'complex', 'integer', 'character', 'logical')
dataStructures <- c('data.frame', 'vector', 'list', 'matrix')
answer <- list(dataTypes, dataStructures)
Note: it’s nice to make a list in big writing on the board or taped to the wall listing all of these types and structures - leave it up for the rest of the workshop to remind people of the importance of these basics.
Challenge 9
Consider the R output of the matrix below:
OUTPUT
[,1] [,2]
[1,] 4 1
[2,] 9 5
[3,] 10 7What was the correct command used to write this matrix? Examine each command and try to figure out the correct one before typing them. Think about what matrices the other commands will produce.
matrix(c(4, 1, 9, 5, 10, 7), nrow = 3)matrix(c(4, 9, 10, 1, 5, 7), ncol = 2, byrow = TRUE)matrix(c(4, 9, 10, 1, 5, 7), nrow = 2)matrix(c(4, 1, 9, 5, 10, 7), ncol = 2, byrow = TRUE)
Consider the R output of the matrix below:
OUTPUT
[,1] [,2]
[1,] 4 1
[2,] 9 5
[3,] 10 7What was the correct command used to write this matrix? Examine each command and try to figure out the correct one before typing them. Think about what matrices the other commands will produce.
R
matrix(c(4, 1, 9, 5, 10, 7), ncol = 2, byrow = TRUE)
- Use
read.csvto read tabular data in R. - The basic data types in R are double, integer, complex, logical, and character.
- Data structures such as data frames or matrices are built on top of lists and vectors, with some added attributes.
Content from Exploring Data Frames
Last updated on 2026-04-21 | Edit this page
Estimated time: 30 minutes
Overview
Questions
- How can I manipulate a data frame?
Objectives
- Add and remove rows or columns.
- Append two data frames.
- Display basic properties of data frames including size and class of the columns, names, and first few rows.
At this point, you’ve seen it all: in the last lesson, we toured all the basic data types and data structures in R. Everything you do will be a manipulation of those tools. But most of the time, the star of the show is the data frame—the table that we created by loading information from a csv file. In this lesson, we’ll learn a few more things about working with data frames.
Adding columns and rows in data frames
We already learned that the columns of a data frame are vectors, so that our data are consistent in type throughout the columns. As such, if we want to add a new column, we can start by making a new vector:
R
age <- c(2, 3, 5)
cats
OUTPUT
coat weight likes_catnip
1 calico 2.1 1
2 black 5.0 0
3 tabby 3.2 1We can then add this as a column via:
R
cbind(cats, age)
OUTPUT
coat weight likes_catnip age
1 calico 2.1 1 2
2 black 5.0 0 3
3 tabby 3.2 1 5Note that if we tried to add a vector of ages with a different number of entries than the number of rows in the data frame, it would fail:
R
age <- c(2, 3, 5, 12)
cbind(cats, age)
ERROR
Error in `data.frame()`:
! arguments imply differing number of rows: 3, 4R
age <- c(2, 3)
cbind(cats, age)
ERROR
Error in `data.frame()`:
! arguments imply differing number of rows: 3, 2Why didn’t this work? Of course, R wants to see one element in our new column for every row in the table:
R
nrow(cats)
OUTPUT
[1] 3R
length(age)
OUTPUT
[1] 2So for it to work we need to have nrow(cats) =
length(age). Let’s overwrite the content of cats with our
new data frame.
R
age <- c(2, 3, 5)
cats <- cbind(cats, age)
Now how about adding rows? We already know that the rows of a data frame are lists:
R
newRow <- list("tortoiseshell", 3.3, TRUE, 9)
cats <- rbind(cats, newRow)
Let’s confirm that our new row was added correctly.
R
cats
OUTPUT
coat weight likes_catnip age
1 calico 2.1 1 2
2 black 5.0 0 3
3 tabby 3.2 1 5
4 tortoiseshell 3.3 1 9Removing rows
We now know how to add rows and columns to our data frame in R. Now let’s learn to remove rows.
R
cats
OUTPUT
coat weight likes_catnip age
1 calico 2.1 1 2
2 black 5.0 0 3
3 tabby 3.2 1 5
4 tortoiseshell 3.3 1 9We can ask for a data frame minus the last row:
R
cats[-4, ]
OUTPUT
coat weight likes_catnip age
1 calico 2.1 1 2
2 black 5.0 0 3
3 tabby 3.2 1 5Notice the comma with nothing after it to indicate that we want to drop the entire fourth row.
Note: we could also remove several rows at once by putting the row
numbers inside of a vector, for example:
cats[c(-3,-4), ]
Removing columns
We can also remove columns in our data frame. What if we want to remove the column “age”. We can remove it in two ways, by variable number or by index.
R
cats[,-4]
OUTPUT
coat weight likes_catnip
1 calico 2.1 1
2 black 5.0 0
3 tabby 3.2 1
4 tortoiseshell 3.3 1Notice the comma with nothing before it, indicating we want to keep all of the rows.
Alternatively, we can drop the column by using the index name and the
%in% operator. The %in% operator goes through
each element of its left argument, in this case the names of
cats, and asks, “Does this element occur in the second
argument?”
R
drop <- names(cats) %in% c("age")
cats[,!drop]
OUTPUT
coat weight likes_catnip
1 calico 2.1 1
2 black 5.0 0
3 tabby 3.2 1
4 tortoiseshell 3.3 1We will cover subsetting with logical operators like
%in% in more detail in the next episode. See the section Subsetting through other logical
operations
Appending to a data frame
The key to remember when adding data to a data frame is that
columns are vectors and rows are lists. We can also glue two
data frames together with rbind:
R
cats <- rbind(cats, cats)
cats
OUTPUT
coat weight likes_catnip age
1 calico 2.1 1 2
2 black 5.0 0 3
3 tabby 3.2 1 5
4 tortoiseshell 3.3 1 9
5 calico 2.1 1 2
6 black 5.0 0 3
7 tabby 3.2 1 5
8 tortoiseshell 3.3 1 9Challenge 1
You can create a new data frame right from within R with the following syntax:
R
df <- data.frame(id = c("a", "b", "c"),
x = 1:3,
y = c(TRUE, TRUE, FALSE))
Make a data frame that holds the following information for yourself:
- first name
- last name
- lucky number
Then use rbind to add an entry for the people sitting
beside you. Finally, use cbind to add a column with each
person’s answer to the question, “Is it time for coffee break?”
R
df <- data.frame(first = c("Grace"),
last = c("Hopper"),
lucky_number = c(0))
df <- rbind(df, list("Marie", "Curie", 238) )
df <- cbind(df, coffeetime = c(TRUE,TRUE))
Realistic example
So far, you have seen the basics of manipulating data frames with our
cat data; now let’s use those skills to digest a more realistic dataset.
Let’s read in the gapminder dataset that we downloaded
previously:
R
gapminder <- read.csv("data/gapminder_data.csv")
Miscellaneous Tips
Another type of file you might encounter are tab-separated value files (.tsv). To specify a tab as a separator, use
"\\t"orread.delim().Files can also be downloaded directly from the Internet into a local folder of your choice onto your computer using the
download.filefunction. Theread.csvfunction can then be executed to read the downloaded file from the download location, for example,
R
download.file("https://raw.githubusercontent.com/swcarpentry/r-novice-gapminder/main/episodes/data/gapminder_data.csv", destfile = "data/gapminder_data.csv")
gapminder <- read.csv("data/gapminder_data.csv")
- Alternatively, you can also read in files directly into R from the
Internet by replacing the file paths with a web address in
read.csv. One should note that in doing this no local copy of the csv file is first saved onto your computer. For example,
R
gapminder <- read.csv("https://raw.githubusercontent.com/swcarpentry/r-novice-gapminder/main/episodes/data/gapminder_data.csv")
You can read directly from excel spreadsheets without converting them to plain text first by using the readxl package.
The argument “stringsAsFactors” can be useful to tell R how to read strings either as factors or as character strings. In R versions after 4.0, all strings are read-in as characters by default, but in earlier versions of R, strings are read-in as factors by default. For more information, see the call-out in the previous episode.
Let’s investigate gapminder a bit; the first thing we should always
do is check out what the data looks like with str:
R
str(gapminder)
OUTPUT
'data.frame': 1704 obs. of 6 variables:
$ country : chr "Afghanistan" "Afghanistan" "Afghanistan" "Afghanistan" ...
$ year : int 1952 1957 1962 1967 1972 1977 1982 1987 1992 1997 ...
$ pop : num 8425333 9240934 10267083 11537966 13079460 ...
$ continent: chr "Asia" "Asia" "Asia" "Asia" ...
$ lifeExp : num 28.8 30.3 32 34 36.1 ...
$ gdpPercap: num 779 821 853 836 740 ...An additional method for examining the structure of gapminder is to
use the summary function. This function can be used on
various objects in R. For data frames, summary yields a
numeric, tabular, or descriptive summary of each column. Numeric or
integer columns are described by the descriptive statistics (quartiles
and mean), and character columns by its length, class, and mode.
R
summary(gapminder)
OUTPUT
country year pop continent
Length:1704 Min. :1952 Min. :6.001e+04 Length:1704
Class :character 1st Qu.:1966 1st Qu.:2.794e+06 Class :character
Mode :character Median :1980 Median :7.024e+06 Mode :character
Mean :1980 Mean :2.960e+07
3rd Qu.:1993 3rd Qu.:1.959e+07
Max. :2007 Max. :1.319e+09
lifeExp gdpPercap
Min. :23.60 Min. : 241.2
1st Qu.:48.20 1st Qu.: 1202.1
Median :60.71 Median : 3531.8
Mean :59.47 Mean : 7215.3
3rd Qu.:70.85 3rd Qu.: 9325.5
Max. :82.60 Max. :113523.1 Along with the str and summary functions,
we can examine individual columns of the data frame with our
typeof function:
R
typeof(gapminder$year)
OUTPUT
[1] "integer"R
typeof(gapminder$country)
OUTPUT
[1] "character"R
str(gapminder$country)
OUTPUT
chr [1:1704] "Afghanistan" "Afghanistan" "Afghanistan" "Afghanistan" ...We can also interrogate the data frame for information about its
dimensions; remembering that str(gapminder) said there were
1704 observations of 6 variables in gapminder, what do you think the
following will produce, and why?
R
length(gapminder)
OUTPUT
[1] 6A fair guess would have been to say that the length of a data frame would be the number of rows it has (1704), but this is not the case; remember, a data frame is a list of vectors and factors:
R
typeof(gapminder)
OUTPUT
[1] "list"When length gave us 6, it’s because gapminder is built
out of a list of 6 columns. To get the number of rows and columns in our
dataset, try:
R
nrow(gapminder)
OUTPUT
[1] 1704R
ncol(gapminder)
OUTPUT
[1] 6Or, both at once:
R
dim(gapminder)
OUTPUT
[1] 1704 6We’ll also likely want to know what the titles of all the columns are, so we can ask for them later:
R
colnames(gapminder)
OUTPUT
[1] "country" "year" "pop" "continent" "lifeExp" "gdpPercap"At this stage, it’s important to ask ourselves if the structure R is reporting matches our intuition or expectations; do the basic data types reported for each column make sense? If not, we need to sort any problems out now before they turn into bad surprises down the road, using what we’ve learned about how R interprets data, and the importance of strict consistency in how we record our data.
Once we’re happy that the data types and structures seem reasonable, it’s time to start digging into our data proper. Check out the first few lines:
R
head(gapminder)
OUTPUT
country year pop continent lifeExp gdpPercap
1 Afghanistan 1952 8425333 Asia 28.801 779.4453
2 Afghanistan 1957 9240934 Asia 30.332 820.8530
3 Afghanistan 1962 10267083 Asia 31.997 853.1007
4 Afghanistan 1967 11537966 Asia 34.020 836.1971
5 Afghanistan 1972 13079460 Asia 36.088 739.9811
6 Afghanistan 1977 14880372 Asia 38.438 786.1134Challenge 2
It’s good practice to also check the last few lines of your data and some in the middle. How would you do this?
Searching for ones specifically in the middle isn’t too hard, but we could ask for a few lines at random. How would you code this?
To check the last few lines it’s relatively simple as R already has a function for this:
R
tail(gapminder)
tail(gapminder, n = 15)
What about a few arbitrary rows just in case something is odd in the middle?
Tip: There are several ways to achieve this.
The solution here presents one form of using nested functions, i.e. a function passed as an argument to another function. This might sound like a new concept, but you are already using it! Remember my_dataframe[rows, cols] will print to screen your data frame with the number of rows and columns you asked for (although you might have asked for a range or named columns for example). How would you get the last row if you don’t know how many rows your data frame has? R has a function for this. What about getting a (pseudorandom) sample? R also has a function for this.
R
gapminder[sample(nrow(gapminder), 5), ]
To make sure our analysis is reproducible, we should put the code into a script file so we can come back to it later.
Challenge 3
Go to file -> new file -> R script, and write an R script to
load in the gapminder dataset. Put it in the scripts/
directory and add it to version control.
Run the script using the source function, using the file
path as its argument (or by pressing the “source” button in
RStudio).
The source function can be used to use a script within a
script. Assume you would like to load the same type of file over and
over again and therefore you need to specify the arguments to fit the
needs of your file. Instead of writing the necessary argument again and
again you could just write it once and save it as a script. Then, you
can use source("Your_Script_containing_the_load_function")
in a new script to use the function of that script without writing
everything again. Check out ?source to find out more.
R
download.file("https://raw.githubusercontent.com/swcarpentry/r-novice-gapminder/main/episodes/data/gapminder_data.csv", destfile = "data/gapminder_data.csv")
gapminder <- read.csv(file = "data/gapminder_data.csv")
To run the script and load the data into the gapminder
variable:
R
source(file = "scripts/load-gapminder.R")
Challenge 4
Read the output of str(gapminder) again; this time, use
what you’ve learned about lists and vectors, as well as the output of
functions like colnames and dim to explain
what everything that str prints out for gapminder means. If
there are any parts you can’t interpret, discuss with your
neighbors!
The object gapminder is a data frame with columns
-
countryandcontinentare character strings. -
yearis an integer vector. -
pop,lifeExp, andgdpPercapare numeric vectors.
- Use
cbind()to add a new column to a data frame. - Use
rbind()to add a new row to a data frame. - Remove rows from a data frame.
- Use
str(),summary(),nrow(),ncol(),dim(),colnames(),head(), andtypeof()to understand the structure of a data frame. - Read in a csv file using
read.csv(). - Understand what
length()of a data frame represents.
Content from Subsetting Data
Last updated on 2026-04-21 | Edit this page
Estimated time: 50 minutes
Overview
Questions
- How can I work with subsets of data in R?
Objectives
- To be able to subset vectors, factors, matrices, lists, and data frames
- To be able to extract individual and multiple elements: by index, by name, using comparison operations
- To be able to skip and remove elements from various data structures.
R has many powerful subset operators. Mastering them will allow you to easily perform complex operations on any kind of dataset.
There are six different ways we can subset any kind of object, and three different subsetting operators for the different data structures.
Let’s start with the workhorse of R: a simple numeric vector.
R
x <- c(5.4, 6.2, 7.1, 4.8, 7.5)
names(x) <- c('a', 'b', 'c', 'd', 'e')
x
OUTPUT
a b c d e
5.4 6.2 7.1 4.8 7.5 Atomic vectors
In R, simple vectors containing character strings, numbers, or logical values are called atomic vectors because they can’t be further simplified.
So now that we’ve created a dummy vector to play with, how do we get at its contents?
Accessing elements using their indices
To extract elements of a vector we can give their corresponding index, starting from one:
R
x[1]
OUTPUT
a
5.4 R
x[4]
OUTPUT
d
4.8 It may look different, but the square brackets operator is a function. For vectors (and matrices), it means “get me the nth element”.
We can ask for multiple elements at once:
R
x[c(1, 3)]
OUTPUT
a c
5.4 7.1 Or slices of the vector:
R
x[1:4]
OUTPUT
a b c d
5.4 6.2 7.1 4.8 the : operator creates a sequence of numbers from the
left element to the right.
R
1:4
OUTPUT
[1] 1 2 3 4R
c(1, 2, 3, 4)
OUTPUT
[1] 1 2 3 4We can ask for the same element multiple times:
R
x[c(1,1,3)]
OUTPUT
a a c
5.4 5.4 7.1 If we ask for an index beyond the length of the vector, R will return a missing value:
R
x[6]
OUTPUT
<NA>
NA This is a vector of length one containing an NA, whose
name is also NA.
If we ask for the 0th element, we get an empty vector:
R
x[0]
OUTPUT
named numeric(0)Vector numbering in R starts at 1
In many programming languages (C and Python, for example), the first element of a vector has an index of 0. In R, the first element is 1.
Skipping and removing elements
If we use a negative number as the index of a vector, R will return every element except for the one specified:
R
x[-2]
OUTPUT
a c d e
5.4 7.1 4.8 7.5 We can skip multiple elements:
R
x[c(-1, -5)] # or x[-c(1,5)]
OUTPUT
b c d
6.2 7.1 4.8 Tip: Order of operations
A common trip up for novices occurs when trying to skip slices of a vector. It’s natural to try to negate a sequence like so:
R
x[-1:3]
This gives a somewhat cryptic error:
ERROR
Error in `x[-1:3]`:
! only 0's may be mixed with negative subscriptsBut remember the order of operations. : is really a
function. It takes its first argument as -1, and its second as 3, so
generates the sequence of numbers: c(-1, 0, 1, 2, 3).
The correct solution is to wrap that function call in brackets, so
that the - operator applies to the result:
R
x[-(1:3)]
OUTPUT
d e
4.8 7.5 To remove elements from a vector, we need to assign the result back into the variable:
R
x <- x[-4]
x
OUTPUT
a b c e
5.4 6.2 7.1 7.5 Challenge 1
Given the following code:
R
x <- c(5.4, 6.2, 7.1, 4.8, 7.5)
names(x) <- c('a', 'b', 'c', 'd', 'e')
print(x)
OUTPUT
a b c d e
5.4 6.2 7.1 4.8 7.5 Come up with at least 2 different commands that will produce the following output:
OUTPUT
b c d
6.2 7.1 4.8 After you find 2 different commands, compare notes with your neighbour. Did you have different strategies?
R
x[2:4]
OUTPUT
b c d
6.2 7.1 4.8 R
x[-c(1,5)]
OUTPUT
b c d
6.2 7.1 4.8 R
x[c(2,3,4)]
OUTPUT
b c d
6.2 7.1 4.8 Subsetting by name
We can extract elements by using their name, instead of extracting by index:
R
x <- c(a=5.4, b=6.2, c=7.1, d=4.8, e=7.5) # we can name a vector 'on the fly'
x[c("a", "c")]
OUTPUT
a c
5.4 7.1 This is usually a much more reliable way to subset objects: the position of various elements can often change when chaining together subsetting operations, but the names will always remain the same!
Subsetting through other logical operations
We can also use any logical vector to subset:
R
x[c(FALSE, FALSE, TRUE, FALSE, TRUE)]
OUTPUT
c e
7.1 7.5 Since comparison operators (e.g. >,
<, ==) evaluate to logical vectors, we can
also use them to succinctly subset vectors: the following statement
gives the same result as the previous one.
R
x[x > 7]
OUTPUT
c e
7.1 7.5 Breaking it down, this statement first evaluates x>7,
generating a logical vector
c(FALSE, FALSE, TRUE, FALSE, TRUE), and then selects the
elements of x corresponding to the TRUE
values.
We can use == to mimic the previous method of indexing
by name (remember you have to use == rather than
= for comparisons):
R
x[names(x) == "a"]
OUTPUT
a
5.4 Tip: Combining logical conditions
We often want to combine multiple logical criteria. For example, we might want to find all the countries that are located in Asia or Europe and have life expectancies within a certain range. Several operations for combining logical vectors exist in R:
-
&, the “logical AND” operator: returnsTRUEif both the left and right areTRUE. -
|, the “logical OR” operator: returnsTRUE, if either the left or right (or both) areTRUE.
You may sometimes see && and ||
instead of & and |. These two-character
operators only look at the first element of each vector and ignore the
remaining elements. In general you should not use the two-character
operators in data analysis; save them for programming, i.e. deciding
whether to execute a statement.
-
!, the “logical NOT” operator: convertsTRUEtoFALSEandFALSEtoTRUE. It can negate a single logical condition (eg!TRUEbecomesFALSE), or a whole vector of conditions(eg!c(TRUE, FALSE)becomesc(FALSE, TRUE)).
Additionally, you can compare the elements within a single vector
using the all function (which returns TRUE if
every element of the vector is TRUE) and the
any function (which returns TRUE if one or
more elements of the vector are TRUE).
Challenge 2
Given the following code:
R
x <- c(5.4, 6.2, 7.1, 4.8, 7.5)
names(x) <- c('a', 'b', 'c', 'd', 'e')
print(x)
OUTPUT
a b c d e
5.4 6.2 7.1 4.8 7.5 Write a subsetting command to return the values in x that are greater than 4 and less than 7.
R
x_subset <- x[x<7 & x>4]
print(x_subset)
OUTPUT
a b d
5.4 6.2 4.8 Tip: Non-unique names
You should be aware that it is possible for multiple elements in a vector to have the same name. (For a data frame, columns can have the same name — although R tries to avoid this — but row names must be unique.) Consider these examples:
R
x <- 1:3
x
OUTPUT
[1] 1 2 3R
names(x) <- c('a', 'a', 'a')
x
OUTPUT
a a a
1 2 3 R
x['a'] # only returns first value
OUTPUT
a
1 R
x[names(x) == 'a'] # returns all three values
OUTPUT
a a a
1 2 3 Tip: Getting help for operators
Remember you can search for help on operators by wrapping them in
quotes: help("%in%") or ?"%in%".
Skipping named elements
Skipping or removing named elements is a little harder. If we try to skip one named element by negating the string, R complains (slightly obscurely) that it doesn’t know how to take the negative of a string:
R
x <- c(a=5.4, b=6.2, c=7.1, d=4.8, e=7.5) # we start again by naming a vector 'on the fly'
x[-"a"]
ERROR
Error in `-"a"`:
! invalid argument to unary operatorHowever, we can use the != (not-equals) operator to
construct a logical vector that will do what we want:
R
x[names(x) != "a"]
OUTPUT
b c d e
6.2 7.1 4.8 7.5 Skipping multiple named indices is a little bit harder still. Suppose
we want to drop the "a" and "c" elements, so
we try this:
R
x[names(x)!=c("a","c")]
WARNING
Warning in names(x) != c("a", "c"): longer object length is not a multiple of
shorter object lengthOUTPUT
b c d e
6.2 7.1 4.8 7.5 R did something, but it gave us a warning that we ought to
pay attention to - and it apparently gave us the wrong answer
(the "c" element is still included in the vector)!
So what does != actually do in this case? That’s an
excellent question.
Recycling
Let’s take a look at the comparison component of this code:
R
names(x) != c("a", "c")
WARNING
Warning in names(x) != c("a", "c"): longer object length is not a multiple of
shorter object lengthOUTPUT
[1] FALSE TRUE TRUE TRUE TRUEWhy does R give TRUE as the third element of this
vector, when names(x)[3] != "c" is obviously false? When
you use !=, R tries to compare each element of the left
argument with the corresponding element of its right argument. What
happens when you compare vectors of different lengths?
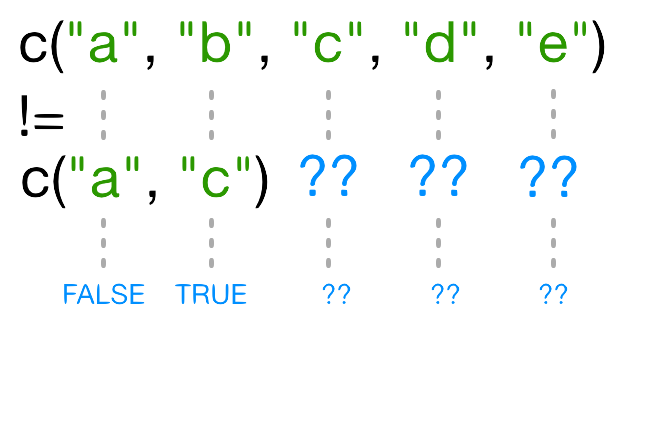
When one vector is shorter than the other, it gets recycled:
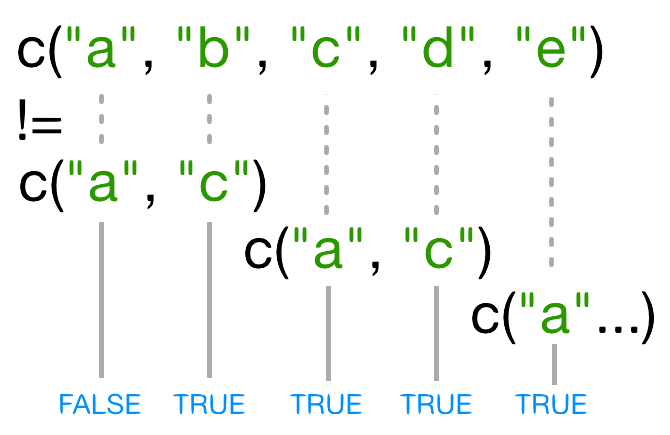
In this case R repeats c("a", "c") as
many times as necessary to match names(x), i.e. we get
c("a","c","a","c","a"). Since the recycled "a"
doesn’t match the third element of names(x), the value of
!= is TRUE. Because in this case the longer
vector length (5) isn’t a multiple of the shorter vector length (2), R
printed a warning message. If we had been unlucky and
names(x) had contained six elements, R would
silently have done the wrong thing (i.e., not what we intended
it to do). This recycling rule can can introduce hard-to-find and subtle
bugs!
The way to get R to do what we really want (match each
element of the left argument with all of the elements of the
right argument) it to use the %in% operator. The
%in% operator goes through each element of its left
argument, in this case the names of x, and asks, “Does this
element occur in the second argument?”. Here, since we want to
exclude values, we also need a ! operator to
change “in” to “not in”:
R
x[! names(x) %in% c("a","c") ]
OUTPUT
b d e
6.2 4.8 7.5 Challenge 3
Selecting elements of a vector that match any of a list of components
is a very common data analysis task. For example, the gapminder data set
contains country and continent variables, but
no information between these two scales. Suppose we want to pull out
information from southeast Asia: how do we set up an operation to
produce a logical vector that is TRUE for all of the
countries in southeast Asia and FALSE otherwise?
Suppose you have these data:
R
seAsia <- c("Myanmar","Thailand","Cambodia","Vietnam","Laos")
## read in the gapminder data that we downloaded in episode 2
gapminder <- read.csv("data/gapminder_data.csv", header=TRUE)
## extract the `country` column from a data frame (we'll see this later);
## convert from a factor to a character;
## and get just the non-repeated elements
countries <- unique(as.character(gapminder$country))
There’s a wrong way (using only ==), which will give you
a warning; a clunky way (using the logical operators == and
|); and an elegant way (using %in%). See
whether you can come up with all three and explain how they (don’t)
work.
- The wrong way to do this problem is
countries==seAsia. This gives a warning ("In countries == seAsia : longer object length is not a multiple of shorter object length") and the wrong answer (a vector of allFALSEvalues), because none of the recycled values ofseAsiahappen to line up correctly with matching values incountry. - The clunky (but technically correct) way to do this problem is
R
(countries=="Myanmar" | countries=="Thailand" |
countries=="Cambodia" | countries == "Vietnam" | countries=="Laos")
(or countries==seAsia[1] | countries==seAsia[2] | ...).
This gives the correct values, but hopefully you can see how awkward it
is (what if we wanted to select countries from a much longer list?).
- The best way to do this problem is
countries %in% seAsia, which is both correct and easy to type (and read).
Handling special values
At some point you will encounter functions in R that cannot handle missing, infinite, or undefined data.
There are a number of special functions you can use to filter out this data:
-
is.nawill return all positions in a vector, matrix, or data.frame containingNA(orNaN) - likewise,
is.nan, andis.infinitewill do the same forNaNandInf. -
is.finitewill return all positions in a vector, matrix, or data.frame that do not containNA,NaNorInf. -
na.omitwill filter out all missing values from a vector
Factor subsetting
Now that we’ve explored the different ways to subset vectors, how do we subset the other data structures?
Factor subsetting works the same way as vector subsetting.
R
f <- factor(c("a", "a", "b", "c", "c", "d"))
f[f == "a"]
OUTPUT
[1] a a
Levels: a b c dR
f[f %in% c("b", "c")]
OUTPUT
[1] b c c
Levels: a b c dR
f[1:3]
OUTPUT
[1] a a b
Levels: a b c dSkipping elements will not remove the level even if no more of that category exists in the factor:
R
f[-3]
OUTPUT
[1] a a c c d
Levels: a b c dMatrix subsetting
Matrices are also subsetted using the [ function. In
this case it takes two arguments: the first applying to the rows, the
second to its columns:
R
set.seed(1)
m <- matrix(rnorm(6*4), ncol=4, nrow=6)
m[3:4, c(3,1)]
OUTPUT
[,1] [,2]
[1,] 1.12493092 -0.8356286
[2,] -0.04493361 1.5952808You can leave the first or second arguments blank to retrieve all the rows or columns respectively:
R
m[, c(3,4)]
OUTPUT
[,1] [,2]
[1,] -0.62124058 0.82122120
[2,] -2.21469989 0.59390132
[3,] 1.12493092 0.91897737
[4,] -0.04493361 0.78213630
[5,] -0.01619026 0.07456498
[6,] 0.94383621 -1.98935170If we only access one row or column, R will automatically convert the result to a vector:
R
m[3,]
OUTPUT
[1] -0.8356286 0.5757814 1.1249309 0.9189774If you want to keep the output as a matrix, you need to specify a
third argument; drop = FALSE:
R
m[3, , drop=FALSE]
OUTPUT
[,1] [,2] [,3] [,4]
[1,] -0.8356286 0.5757814 1.124931 0.9189774Unlike vectors, if we try to access a row or column outside of the matrix, R will throw an error:
R
m[, c(3,6)]
ERROR
Error in `m[, c(3, 6)]`:
! subscript out of boundsTip: Higher dimensional arrays
when dealing with multi-dimensional arrays, each argument to
[ corresponds to a dimension. For example, a 3D array, the
first three arguments correspond to the rows, columns, and depth
dimension.
Because matrices are vectors, we can also subset using only one argument:
R
m[5]
OUTPUT
[1] 0.3295078This usually isn’t useful, and often confusing to read. However it is useful to note that matrices are laid out in column-major format by default. That is the elements of the vector are arranged column-wise:
R
matrix(1:6, nrow=2, ncol=3)
OUTPUT
[,1] [,2] [,3]
[1,] 1 3 5
[2,] 2 4 6If you wish to populate the matrix by row, use
byrow=TRUE:
R
matrix(1:6, nrow=2, ncol=3, byrow=TRUE)
OUTPUT
[,1] [,2] [,3]
[1,] 1 2 3
[2,] 4 5 6Matrices can also be subsetted using their rownames and column names instead of their row and column indices.
Challenge 4
Given the following code:
R
m <- matrix(1:18, nrow=3, ncol=6)
print(m)
OUTPUT
[,1] [,2] [,3] [,4] [,5] [,6]
[1,] 1 4 7 10 13 16
[2,] 2 5 8 11 14 17
[3,] 3 6 9 12 15 18- Which of the following commands will extract the values 11 and 14?
A. m[2,4,2,5]
B. m[2:5]
C. m[4:5,2]
D. m[2,c(4,5)]
D
List subsetting
Now we’ll introduce some new subsetting operators. There are three
functions used to subset lists. We’ve already seen these when learning
about atomic vectors and matrices: [, [[, and
$.
Using [ will always return a list. If you want to
subset a list, but not extract an element, then you
will likely use [.
R
xlist <- list(a = "Software Carpentry", b = 1:10, data = head(mtcars))
xlist[1]
OUTPUT
$a
[1] "Software Carpentry"This returns a list with one element.
We can subset elements of a list exactly the same way as atomic
vectors using [. Comparison operations however won’t work
as they’re not recursive, they will try to condition on the data
structures in each element of the list, not the individual elements
within those data structures.
R
xlist[1:2]
OUTPUT
$a
[1] "Software Carpentry"
$b
[1] 1 2 3 4 5 6 7 8 9 10To extract individual elements of a list, you need to use the
double-square bracket function: [[.
R
xlist[[1]]
OUTPUT
[1] "Software Carpentry"Notice that now the result is a vector, not a list.
You can’t extract more than one element at once:
R
xlist[[1:2]]
ERROR
Error in `xlist[[1:2]]`:
! subscript out of boundsNor use it to skip elements:
R
xlist[[-1]]
ERROR
Error in `xlist[[-1]]`:
! invalid negative subscript in get1index <real>But you can use names to both subset and extract elements:
R
xlist[["a"]]
OUTPUT
[1] "Software Carpentry"The $ function is a shorthand way for extracting
elements by name:
R
xlist$data
OUTPUT
mpg cyl disp hp drat wt qsec vs am gear carb
Mazda RX4 21.0 6 160 110 3.90 2.620 16.46 0 1 4 4
Mazda RX4 Wag 21.0 6 160 110 3.90 2.875 17.02 0 1 4 4
Datsun 710 22.8 4 108 93 3.85 2.320 18.61 1 1 4 1
Hornet 4 Drive 21.4 6 258 110 3.08 3.215 19.44 1 0 3 1
Hornet Sportabout 18.7 8 360 175 3.15 3.440 17.02 0 0 3 2
Valiant 18.1 6 225 105 2.76 3.460 20.22 1 0 3 1Challenge 5
Given the following list:
R
xlist <- list(a = "Software Carpentry", b = 1:10, data = head(mtcars))
Using your knowledge of both list and vector subsetting, extract the number 2 from xlist. Hint: the number 2 is contained within the “b” item in the list.
R
xlist$b[2]
OUTPUT
[1] 2R
xlist[[2]][2]
OUTPUT
[1] 2R
xlist[["b"]][2]
OUTPUT
[1] 2Challenge 6
Given a linear model:
R
mod <- aov(pop ~ lifeExp, data=gapminder)
Extract the residual degrees of freedom (hint:
attributes() will help you)
R
attributes(mod) ## `df.residual` is one of the names of `mod`
R
mod$df.residual
Data frames
Remember the data frames are lists underneath the hood, so similar rules apply. However they are also two dimensional objects:
[ with one argument will act the same way as for lists,
where each list element corresponds to a column. The resulting object
will be a data frame:
R
head(gapminder[3])
OUTPUT
pop
1 8425333
2 9240934
3 10267083
4 11537966
5 13079460
6 14880372Similarly, [[ will act to extract a single
column:
R
head(gapminder[["lifeExp"]])
OUTPUT
[1] 28.801 30.332 31.997 34.020 36.088 38.438And $ provides a convenient shorthand to extract columns
by name:
R
head(gapminder$year)
OUTPUT
[1] 1952 1957 1962 1967 1972 1977With two arguments, [ behaves the same way as for
matrices:
R
gapminder[1:3,]
OUTPUT
country year pop continent lifeExp gdpPercap
1 Afghanistan 1952 8425333 Asia 28.801 779.4453
2 Afghanistan 1957 9240934 Asia 30.332 820.8530
3 Afghanistan 1962 10267083 Asia 31.997 853.1007If we subset a single row, the result will be a data frame (because the elements are mixed types):
R
gapminder[3,]
OUTPUT
country year pop continent lifeExp gdpPercap
3 Afghanistan 1962 10267083 Asia 31.997 853.1007But for a single column the result will be a vector (this can be
changed with the third argument, drop = FALSE).
Challenge 7
Fix each of the following common data frame subsetting errors:
- Extract observations collected for the year 1957
- Extract all columns except 1 through to 4
R
gapminder[,-1:4]
- Extract the rows where the life expectancy is longer the 80 years
R
gapminder[gapminder$lifeExp > 80]
- Extract the first row, and the fourth and fifth columns
(
continentandlifeExp).
R
gapminder[1, 4, 5]
- Advanced: extract rows that contain information for the years 2002 and 2007
R
gapminder[gapminder$year == 2002 | 2007,]
Fix each of the following common data frame subsetting errors:
- Extract observations collected for the year 1957
R
# gapminder[gapminder$year = 1957,]
gapminder[gapminder$year == 1957,]
- Extract all columns except 1 through to 4
R
# gapminder[,-1:4]
gapminder[,-c(1:4)]
- Extract the rows where the life expectancy is longer than 80 years
R
# gapminder[gapminder$lifeExp > 80]
gapminder[gapminder$lifeExp > 80,]
- Extract the first row, and the fourth and fifth columns
(
continentandlifeExp).
R
# gapminder[1, 4, 5]
gapminder[1, c(4, 5)]
- Advanced: extract rows that contain information for the years 2002 and 2007
R
# gapminder[gapminder$year == 2002 | 2007,]
gapminder[gapminder$year == 2002 | gapminder$year == 2007,]
gapminder[gapminder$year %in% c(2002, 2007),]
Challenge 8
Why does
gapminder[1:20]return an error? How does it differ fromgapminder[1:20, ]?Create a new
data.framecalledgapminder_smallthat only contains rows 1 through 9 and 19 through 23. You can do this in one or two steps.
gapminderis a data.frame so needs to be subsetted on two dimensions.gapminder[1:20, ]subsets the data to give the first 20 rows and all columns.
R
gapminder_small <- gapminder[c(1:9, 19:23),]
- Indexing in R starts at 1, not 0.
- Access individual values by location using
[]. - Access slices of data using
[low:high]. - Access arbitrary sets of data using
[c(...)]. - Use logical operations and logical vectors to access subsets of data.
Content from Control Flow
Last updated on 2026-04-21 | Edit this page
Estimated time: 65 minutes
Overview
Questions
- How can I make data-dependent choices in R?
- How can I repeat operations in R?
Objectives
- Write conditional statements with
if...elsestatements andifelse(). - Write and understand
for()loops.
Often when we’re coding we want to control the flow of our actions. This can be done by setting actions to occur only if a condition or a set of conditions are met. Alternatively, we can also set an action to occur a particular number of times.
There are several ways you can control flow in R. For conditional statements, the most commonly used approaches are the constructs:
R
# if
if (condition is true) {
perform action
}
# if ... else
if (condition is true) {
perform action
} else { # that is, if the condition is false,
perform alternative action
}Say, for example, that we want R to print a message if a variable
x has a particular value:
R
x <- 8
if (x >= 10) {
print("x is greater than or equal to 10")
}
x
OUTPUT
[1] 8The print statement does not appear in the console because x is not
greater than 10. To print a different message for numbers less than 10,
we can add an else statement.
R
x <- 8
if (x >= 10) {
print("x is greater than or equal to 10")
} else {
print("x is less than 10")
}
OUTPUT
[1] "x is less than 10"You can also test multiple conditions by using
else if.
R
x <- 8
if (x >= 10) {
print("x is greater than or equal to 10")
} else if (x > 5) {
print("x is greater than 5, but less than 10")
} else {
print("x is less than 5")
}
OUTPUT
[1] "x is greater than 5, but less than 10"Important: when R evaluates the condition inside
if() statements, it is looking for a logical element, i.e.,
TRUE or FALSE. This can cause some headaches
for beginners. For example:
R
x <- 4 == 3
if (x) {
"4 equals 3"
} else {
"4 does not equal 3"
}
OUTPUT
[1] "4 does not equal 3"As we can see, the not equal message was printed because the vector x
is FALSE
R
x <- 4 == 3
x
OUTPUT
[1] FALSEChallenge 1
Use an if() statement to print a suitable message
reporting whether there are any records from 2002 in the
gapminder dataset. Now do the same for 2012.
We will first see a solution to Challenge 1 which does not use the
any() function. We first obtain a logical vector describing
which element of gapminder$year is equal to
2002:
R
gapminder[(gapminder$year == 2002),]
Then, we count the number of rows of the data.frame
gapminder that correspond to the 2002:
R
rows2002_number <- nrow(gapminder[(gapminder$year == 2002),])
The presence of any record for the year 2002 is equivalent to the
request that rows2002_number is one or more:
R
rows2002_number >= 1
Putting all together, we obtain:
R
if(nrow(gapminder[(gapminder$year == 2002),]) >= 1){
print("Record(s) for the year 2002 found.")
}
All this can be done more quickly with any(). The
logical condition can be expressed as:
R
if(any(gapminder$year == 2002)){
print("Record(s) for the year 2002 found.")
}
Did anyone get a warning message like this?
ERROR
Error in `if (gapminder$year == 2012) ...`:
! the condition has length > 1The if() function only accepts singular (of length 1)
inputs, and therefore returns an error when you use it with a vector.
The if() function will still run, but will only evaluate
the condition in the first element of the vector. Therefore, to use the
if() function, you need to make sure your input is singular
(of length 1).
Tip: Built in ifelse()
function
R accepts both if() and
else if() statements structured as outlined above, but also
statements using R’s built-in ifelse()
function. This function accepts both singular and vector inputs and is
structured as follows:
where the first argument is the condition or a set of conditions to
be met, the second argument is the statement that is evaluated when the
condition is TRUE, and the third statement is the statement
that is evaluated when the condition is FALSE.
R
y <- -3
ifelse(y < 0, "y is a negative number", "y is either positive or zero")
OUTPUT
[1] "y is a negative number"Tip: any() and
all()
The any() function will return TRUE if at
least one TRUE value is found within a vector, otherwise it
will return FALSE. This can be used in a similar way to the
%in% operator. The function all(), as the name
suggests, will only return TRUE if all values in the vector
are TRUE.
Repeating operations
If you want to iterate over a set of values, when the order of
iteration is important, and perform the same operation on each, a
for() loop will do the job. We saw for() loops
in the shell
lessons earlier. This is the most flexible of looping operations,
but therefore also the hardest to use correctly. In general, the advice
of many R users would be to learn about for()
loops, but to avoid using for() loops unless the order of
iteration is important: i.e. the calculation at each iteration depends
on the results of previous iterations. If the order of iteration is not
important, then you should learn about vectorized alternatives, such as
the purrr package, as they pay off in computational
efficiency.
The basic structure of a for() loop is:
For example:
R
for (i in 1:10) {
print(i)
}
OUTPUT
[1] 1
[1] 2
[1] 3
[1] 4
[1] 5
[1] 6
[1] 7
[1] 8
[1] 9
[1] 10The 1:10 bit creates a vector on the fly; you can
iterate over any other vector as well.
We can use a for() loop nested within another
for() loop to iterate over two things at once.
R
for (i in 1:5) {
for (j in c('a', 'b', 'c', 'd', 'e')) {
print(paste(i,j))
}
}
OUTPUT
[1] "1 a"
[1] "1 b"
[1] "1 c"
[1] "1 d"
[1] "1 e"
[1] "2 a"
[1] "2 b"
[1] "2 c"
[1] "2 d"
[1] "2 e"
[1] "3 a"
[1] "3 b"
[1] "3 c"
[1] "3 d"
[1] "3 e"
[1] "4 a"
[1] "4 b"
[1] "4 c"
[1] "4 d"
[1] "4 e"
[1] "5 a"
[1] "5 b"
[1] "5 c"
[1] "5 d"
[1] "5 e"We notice in the output that when the first index (i) is
set to 1, the second index (j) iterates through its full
set of indices. Once the indices of j have been iterated
through, then i is incremented. This process continues
until the last index has been used for each for() loop.
Rather than printing the results, we could write the loop output to a new object.
R
output_vector <- c()
for (i in 1:5) {
for (j in c('a', 'b', 'c', 'd', 'e')) {
temp_output <- paste(i, j)
output_vector <- c(output_vector, temp_output)
}
}
output_vector
OUTPUT
[1] "1 a" "1 b" "1 c" "1 d" "1 e" "2 a" "2 b" "2 c" "2 d" "2 e" "3 a" "3 b"
[13] "3 c" "3 d" "3 e" "4 a" "4 b" "4 c" "4 d" "4 e" "5 a" "5 b" "5 c" "5 d"
[25] "5 e"This approach can be useful, but ‘growing your results’ (building the result object incrementally) is computationally inefficient, so avoid it when you are iterating through a lot of values.
Tip: don’t grow your results
One of the biggest things that trips up novices and experienced R users alike, is building a results object (vector, list, matrix, data frame) as your for loop progresses. Computers are very bad at handling this, so your calculations can very quickly slow to a crawl. It’s much better to define an empty results object before hand of appropriate dimensions, rather than initializing an empty object without dimensions. So if you know the end result will be stored in a matrix like above, create an empty matrix with 5 row and 5 columns, then at each iteration store the results in the appropriate location.
A better way is to define your (empty) output object before filling in the values. For this example, it looks more involved, but is still more efficient.
R
output_matrix <- matrix(nrow = 5, ncol = 5)
j_vector <- c('a', 'b', 'c', 'd', 'e')
for (i in 1:5) {
for (j in 1:5) {
temp_j_value <- j_vector[j]
temp_output <- paste(i, temp_j_value)
output_matrix[i, j] <- temp_output
}
}
output_vector2 <- as.vector(output_matrix)
output_vector2
OUTPUT
[1] "1 a" "2 a" "3 a" "4 a" "5 a" "1 b" "2 b" "3 b" "4 b" "5 b" "1 c" "2 c"
[13] "3 c" "4 c" "5 c" "1 d" "2 d" "3 d" "4 d" "5 d" "1 e" "2 e" "3 e" "4 e"
[25] "5 e"Tip: While loops
Sometimes you will find yourself needing to repeat an operation as
long as a certain condition is met. You can do this with a
while() loop.
R will interpret a condition being met as “TRUE”.
As an example, here’s a while loop that generates random numbers from
a uniform distribution (the runif() function) between 0 and
1 until it gets one that’s less than 0.1.
R
z <- 1
while(z > 0.1){
z <- runif(1)
cat(z, "\n")
}
while() loops will not always be appropriate. You have
to be particularly careful that you don’t end up stuck in an infinite
loop because your condition is always met and hence the while statement
never terminates.
Challenge 2
Compare the objects output_vector and
output_vector2. Are they the same? If not, why not? How
would you change the last block of code to make
output_vector2 the same as output_vector?
We can check whether the two vectors are identical using the
all() function:
R
all(output_vector == output_vector2)
However, all the elements of output_vector can be found
in output_vector2:
R
all(output_vector %in% output_vector2)
and vice versa:
R
all(output_vector2 %in% output_vector)
therefore, the element in output_vector and
output_vector2 are just sorted in a different order. This
is because as.vector() outputs the elements of an input
matrix going over its column. Taking a look at
output_matrix, we can notice that we want its elements by
rows. The solution is to transpose the output_matrix. We
can do it either by calling the transpose function t() or
by inputting the elements in the right order. The first solution
requires to change the original
R
output_vector2 <- as.vector(output_matrix)
into
R
output_vector2 <- as.vector(t(output_matrix))
The second solution requires to change
R
output_matrix[i, j] <- temp_output
into
R
output_matrix[j, i] <- temp_output
Challenge 3
Write a script that loops through the gapminder data by
continent and prints out whether the mean life expectancy is smaller or
larger than 50 years.
Step 1: We want to make sure we can extract all the unique values of the continent vector
R
gapminder <- read.csv("data/gapminder_data.csv")
unique(gapminder$continent)
Step 2: We also need to loop over each of these
continents and calculate the average life expectancy for each
subset of data. We can do that as follows:
- Loop over each of the unique values of ‘continent’
- For each value of continent, create a temporary variable storing that subset
- Return the calculated life expectancy to the user by printing the output:
R
for (iContinent in unique(gapminder$continent)) {
tmp <- gapminder[gapminder$continent == iContinent, ]
cat(iContinent, mean(tmp$lifeExp, na.rm = TRUE), "\n")
rm(tmp)
}
Step 3: The exercise only wants the output printed
if the average life expectancy is less than 50 or greater than 50. So we
need to add an if() condition before printing, which
evaluates whether the calculated average life expectancy is above or
below a threshold, and prints an output conditional on the result. We
need to amend (3) from above:
3a. If the calculated life expectancy is less than some threshold (50 years), return the continent and a statement that life expectancy is less than threshold, otherwise return the continent and a statement that life expectancy is greater than threshold:
R
thresholdValue <- 50
for (iContinent in unique(gapminder$continent)) {
tmp <- mean(gapminder[gapminder$continent == iContinent, "lifeExp"])
if (tmp < thresholdValue){
cat("Average Life Expectancy in", iContinent, "is less than", thresholdValue, "\n")
} else {
cat("Average Life Expectancy in", iContinent, "is greater than", thresholdValue, "\n")
} # end if else condition
rm(tmp)
} # end for loop
Challenge 4
Modify the script from Challenge 3 to loop over each country. This time print out whether the life expectancy is smaller than 50, between 50 and 70, or greater than 70.
We modify our solution to Challenge 3 by now adding two thresholds,
lowerThreshold and upperThreshold and
extending our if-else statements:
R
lowerThreshold <- 50
upperThreshold <- 70
for (iCountry in unique(gapminder$country)) {
tmp <- mean(gapminder[gapminder$country == iCountry, "lifeExp"])
if(tmp < lowerThreshold) {
cat("Average Life Expectancy in", iCountry, "is less than", lowerThreshold, "\n")
} else if(tmp > lowerThreshold && tmp < upperThreshold) {
cat("Average Life Expectancy in", iCountry, "is between", lowerThreshold, "and", upperThreshold, "\n")
} else {
cat("Average Life Expectancy in", iCountry, "is greater than", upperThreshold, "\n")
}
rm(tmp)
}
Challenge 5 - Advanced
Write a script that loops over each country in the
gapminder dataset, tests whether the country starts with a
‘B’, and graphs life expectancy against time as a line graph if the mean
life expectancy is under 50 years.
We will use the grep() command that was introduced in
the Unix
Shell lesson to find countries that start with “B.” Lets understand
how to do this first. Following from the Unix shell section we may be
tempted to try the following
R
grep("^B", unique(gapminder$country))
But when we evaluate this command it returns the indices of the
factor variable country that start with “B.” To get the
values, we must add the value=TRUE option to the
grep() command:
R
grep("^B", unique(gapminder$country), value = TRUE)
We will now store these countries in a variable called
candidateCountries, and then loop over each entry in the variable.
Inside the loop, we evaluate the average life expectancy for each
country, and if the average life expectancy is less than 50 we use
base-plot to plot the evolution of average life expectancy using
with() and subset():
R
thresholdValue <- 50
candidateCountries <- grep("^B", unique(gapminder$country), value = TRUE)
for (iCountry in candidateCountries) {
tmp <- mean(gapminder[gapminder$country == iCountry, "lifeExp"])
if (tmp < thresholdValue) {
cat("Average Life Expectancy in", iCountry, "is less than", thresholdValue, "plotting life expectancy graph... \n")
with(subset(gapminder, country == iCountry),
plot(year, lifeExp,
type = "o",
main = paste("Life Expectancy in", iCountry, "over time"),
ylab = "Life Expectancy",
xlab = "Year"
) # end plot
) # end with
} # end if
rm(tmp)
} # end for loop
- Use
ifandelseto make choices. - Use
forto repeat operations.
Content from Creating Publication-Quality Graphics with ggplot2
Last updated on 2026-04-21 | Edit this page
Estimated time: 80 minutes
Overview
Questions
- How can I create publication-quality graphics in R?
Objectives
- To be able to use ggplot2 to generate publication-quality graphics.
- To apply geometry, aesthetic, and statistics layers to a ggplot plot.
- To manipulate the aesthetics of a plot using different colors, shapes, and lines.
- To improve data visualization through transforming scales and paneling by group.
- To save a plot created with ggplot to disk.
Plotting our data is one of the best ways to quickly explore it and the various relationships between variables.
There are three main plotting systems in R, the base plotting system, the lattice package, and the ggplot2 package.
Today we’ll be learning about the ggplot2 package, because it is the most effective for creating publication-quality graphics.
ggplot2 is built on the grammar of graphics, the idea that any plot can be built from the same set of components: a data set, mapping aesthetics, and graphical layers:
Data sets are the data that you, the user, provide.
Mapping aesthetics are what connect the data to the graphics. They tell ggplot2 how to use your data to affect how the graph looks, such as changing what is plotted on the X or Y axis, or the size or color of different data points.
Layers are the actual graphical output from ggplot2. Layers determine what kinds of plot are shown (scatterplot, histogram, etc.), the coordinate system used (rectangular, polar, others), and other important aspects of the plot. The idea of layers of graphics may be familiar to you if you have used image editing programs like Photoshop, Illustrator, or Inkscape.
Let’s start off building an example using the gapminder data from
earlier. The most basic function is ggplot, which lets R
know that we’re creating a new plot. Any of the arguments we give the
ggplot function are the global options for the
plot: they apply to all layers on the plot.
R
library("ggplot2")
ggplot(data = gapminder)

Here we called ggplot and told it what data we want to
show on our figure. This is not enough information for
ggplot to actually draw anything. It only creates a blank
slate for other elements to be added to.
Now we’re going to add in the mapping aesthetics
using the aes function. aes tells
ggplot how variables in the data map to
aesthetic properties of the figure, such as which columns of
the data should be used for the x and
y locations.
R
ggplot(data = gapminder, mapping = aes(x = gdpPercap, y = lifeExp))
Here we told ggplot we want to plot the “gdpPercap”
column of the gapminder data frame on the x-axis, and the “lifeExp”
column on the y-axis. Notice that we didn’t need to explicitly pass
aes these columns
(e.g. x = gapminder[, "gdpPercap"]), this is because
ggplot is smart enough to know to look in the
data for that column!
The final part of making our plot is to tell ggplot how
we want to visually represent the data. We do this by adding a new
layer to the plot using one of the
geom functions.
R
ggplot(data = gapminder, mapping = aes(x = gdpPercap, y = lifeExp)) +
geom_point()
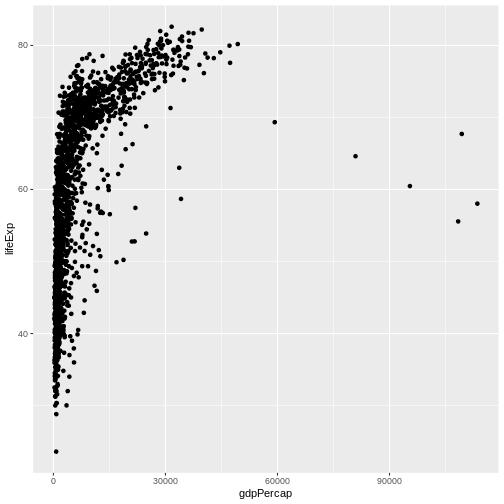
Here we used geom_point, which tells ggplot
we want to visually represent the relationship between
x and y as a scatterplot of
points.
Challenge 1
Modify the example so that the figure shows how life expectancy has changed over time:
R
ggplot(data = gapminder, mapping = aes(x = gdpPercap, y = lifeExp)) + geom_point()
Hint: the gapminder dataset has a column called “year”, which should appear on the x-axis.
Here is one possible solution:
R
ggplot(data = gapminder, mapping = aes(x = year, y = lifeExp)) + geom_point()
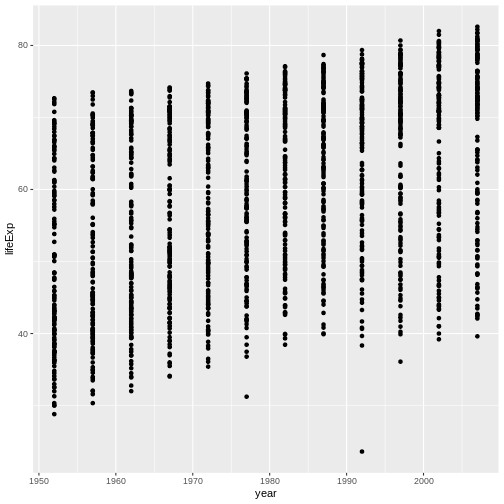
Challenge 2
In the previous examples and challenge we’ve used the
aes function to tell the scatterplot geom
about the x and y locations of each
point. Another aesthetic property we can modify is the point
color. Modify the code from the previous challenge to
color the points by the “continent” column. What trends
do you see in the data? Are they what you expected?
The solution presented below adds color=continent to the
call of the aes function. The general trend seems to
indicate an increased life expectancy over the years. On continents with
stronger economies we find a longer life expectancy.
R
ggplot(data = gapminder, mapping = aes(x = year, y = lifeExp, color=continent)) +
geom_point()
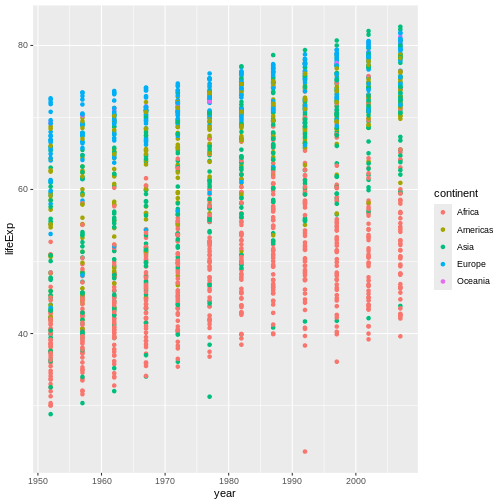
Layers
Using a scatterplot probably isn’t the best for visualizing change
over time. Instead, let’s tell ggplot to visualize the data
as a line plot:
R
ggplot(data = gapminder, mapping = aes(x=year, y=lifeExp, color=continent)) +
geom_line()
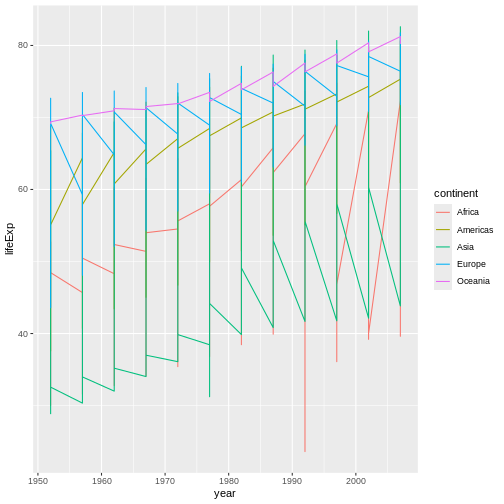
Instead of adding a geom_point layer, we’ve added a
geom_line layer.
However, the result doesn’t look quite as we might have expected: it seems to be jumping around a lot in each continent. Let’s try to separate the data by country, plotting one line for each country:
R
ggplot(data = gapminder, mapping = aes(x=year, y=lifeExp, group=country, color=continent)) +
geom_line()
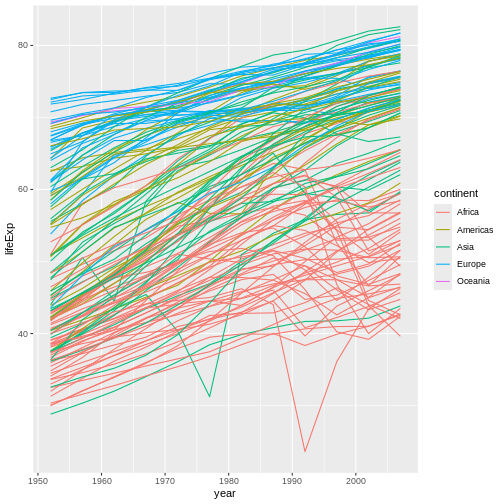
We’ve added the group aesthetic, which
tells ggplot to draw a line for each country.
But what if we want to visualize both lines and points on the plot? We can add another layer to the plot:
R
ggplot(data = gapminder, mapping = aes(x=year, y=lifeExp, group=country, color=continent)) +
geom_line() + geom_point()
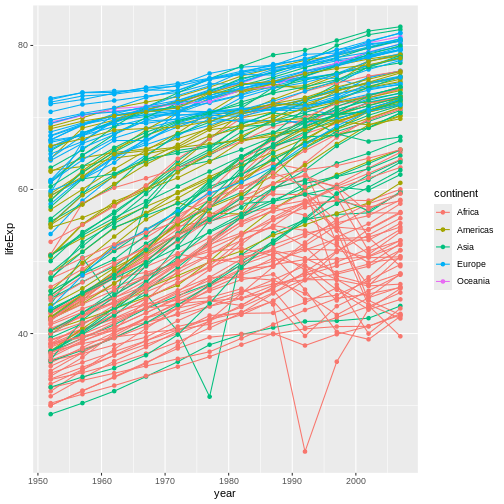
It’s important to note that each layer is drawn on top of the previous layer. In this example, the points have been drawn on top of the lines. Here’s a demonstration:
R
ggplot(data = gapminder, mapping = aes(x=year, y=lifeExp, group=country)) +
geom_line(mapping = aes(color=continent)) + geom_point()
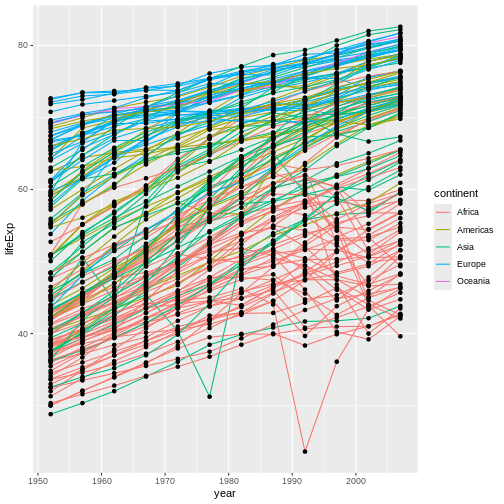
In this example, the aesthetic mapping of
color has been moved from the global plot options in
ggplot to the geom_line layer so it no longer
applies to the points. Now we can clearly see that the points are drawn
on top of the lines.
Tip: Setting an aesthetic to a value instead of a mapping
So far, we’ve seen how to use an aesthetic (such as
color) as a mapping to a variable in the data.
For example, when we use
geom_line(mapping = aes(color=continent)), ggplot will give
a different color to each continent. But what if we want to change the
color of all lines to blue? You may think that
geom_line(mapping = aes(color="blue")) should work, but it
doesn’t. Since we don’t want to create a mapping to a specific variable,
we can move the color specification outside of the aes()
function, like this: geom_line(color="blue").
Challenge 3
Switch the order of the point and line layers from the previous example. What happened?
The lines now get drawn over the points!
R
ggplot(data = gapminder, mapping = aes(x=year, y=lifeExp, group=country)) +
geom_point() + geom_line(mapping = aes(color=continent))
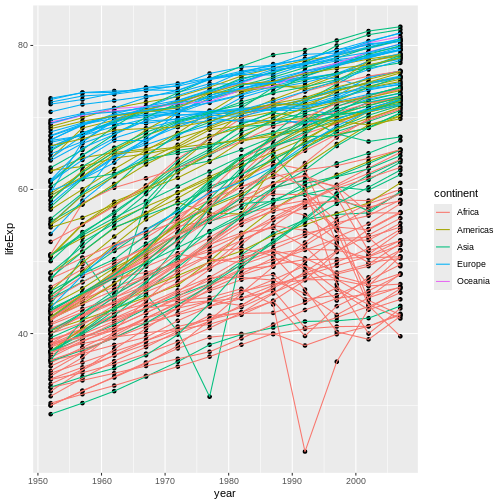
Transformations and statistics
ggplot2 also makes it easy to overlay statistical models over the data. To demonstrate we’ll go back to our first example:
R
ggplot(data = gapminder, mapping = aes(x = gdpPercap, y = lifeExp)) +
geom_point()
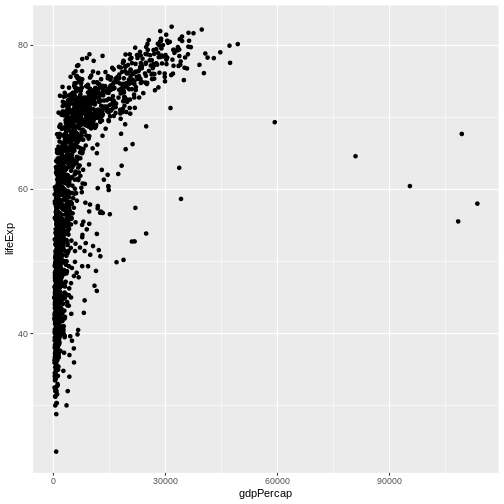
Currently it’s hard to see the relationship between the points due to some strong outliers in GDP per capita. We can change the scale of units on the x axis using the scale functions. These control the mapping between the data values and visual values of an aesthetic. We can also modify the transparency of the points, using the alpha function, which is especially helpful when you have a large amount of data which is very clustered.
R
ggplot(data = gapminder, mapping = aes(x = gdpPercap, y = lifeExp)) +
geom_point(alpha = 0.5) + scale_x_log10()
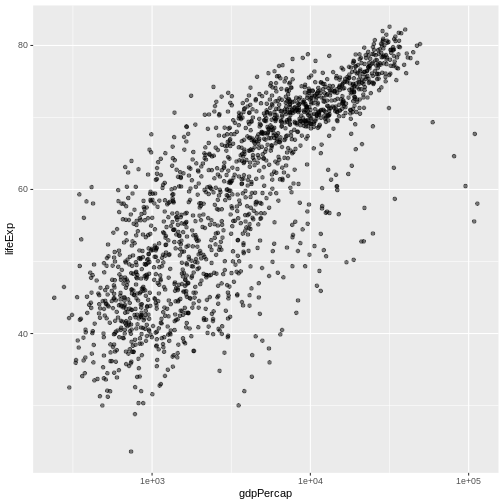
The scale_x_log10 function applied a transformation to
the coordinate system of the plot, so that each multiple of 10 is evenly
spaced from left to right. For example, a GDP per capita of 1,000 is the
same horizontal distance away from a value of 10,000 as the 10,000 value
is from 100,000. This helps to visualize the spread of the data along
the x-axis.
Tip Reminder: Setting an aesthetic to a value instead of a mapping
Notice that we used geom_point(alpha = 0.5). As the
previous tip mentioned, using a setting outside of the
aes() function will cause this value to be used for all
points, which is what we want in this case. But just like any other
aesthetic setting, alpha can also be mapped to a variable in
the data. For example, we can give a different transparency to each
continent with
geom_point(mapping = aes(alpha = continent)).
We can fit a simple relationship to the data by adding another layer,
geom_smooth:
R
ggplot(data = gapminder, mapping = aes(x = gdpPercap, y = lifeExp)) +
geom_point(alpha = 0.5) + scale_x_log10() + geom_smooth(method="lm")
OUTPUT
`geom_smooth()` using formula = 'y ~ x'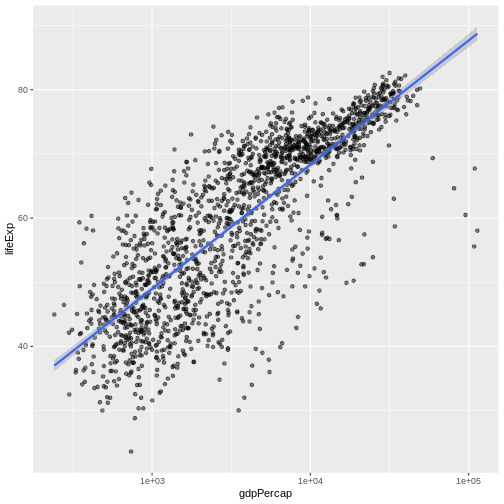
We can make the line thicker by setting the
linewidth aesthetic in the geom_smooth
layer:
R
ggplot(data = gapminder, mapping = aes(x = gdpPercap, y = lifeExp)) +
geom_point(alpha = 0.5) + scale_x_log10() + geom_smooth(method="lm", linewidth=1.5)
OUTPUT
`geom_smooth()` using formula = 'y ~ x'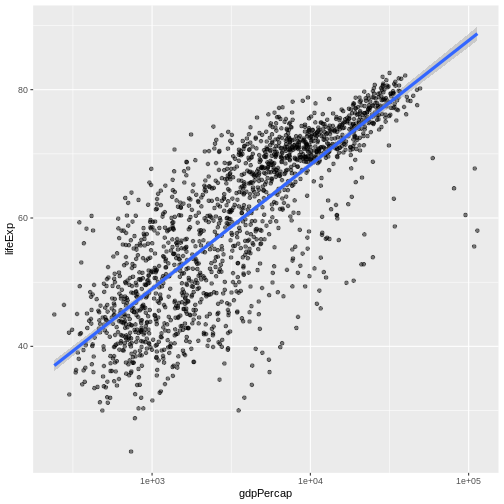
There are two ways an aesthetic can be specified. Here we
set the linewidth aesthetic by passing it as
an argument to geom_smooth and it is applied the same to
the whole geom. Previously in the lesson we’ve used the
aes function to define a mapping between data
variables and their visual representation.
Challenge 4a
Modify the color and size of the points on the point layer in the previous example.
Hint: do not use the aes function.
Hint: the equivalent of linewidth for points is
size.
Here a possible solution: Notice that the color argument
is supplied outside of the aes() function. This means that
it applies to all data points on the graph and is not related to a
specific variable.
R
ggplot(data = gapminder, mapping = aes(x = gdpPercap, y = lifeExp)) +
geom_point(size=3, color="orange") + scale_x_log10() +
geom_smooth(method="lm", linewidth=1.5)
OUTPUT
`geom_smooth()` using formula = 'y ~ x'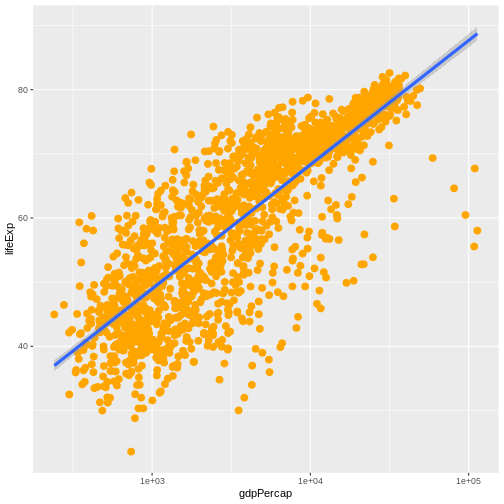
Challenge 4b
Modify your solution to Challenge 4a so that the points are now a different shape and are colored by continent with new trendlines. Hint: The color argument can be used inside the aesthetic.
Here is a possible solution: Notice that supplying the
color argument inside the aes() functions
enables you to connect it to a certain variable. The shape
argument, as you can see, modifies all data points the same way (it is
outside the aes() call) while the color
argument which is placed inside the aes() call modifies a
point’s color based on its continent value.
R
ggplot(data = gapminder, mapping = aes(x = gdpPercap, y = lifeExp, color = continent)) +
geom_point(size=3, shape=17) + scale_x_log10() +
geom_smooth(method="lm", linewidth=1.5)
OUTPUT
`geom_smooth()` using formula = 'y ~ x'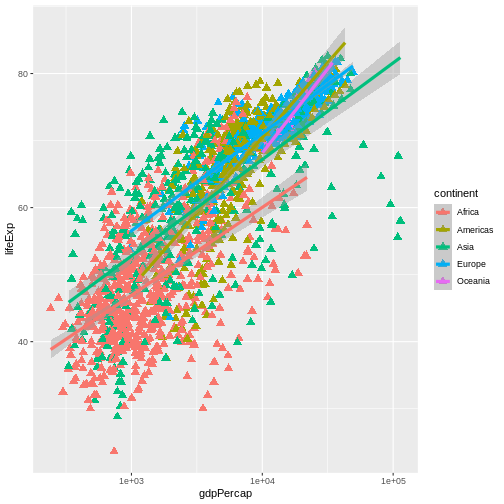
Multi-panel figures
Earlier we visualized the change in life expectancy over time across all countries in one plot. Alternatively, we can split this out over multiple panels by adding a layer of facet panels.
Tip
We start by making a subset of data including only countries located in the Americas. This includes 25 countries, which will begin to clutter the figure. Note that we apply a “theme” definition to rotate the x-axis labels to maintain readability. Nearly everything in ggplot2 is customizable.
R
americas <- gapminder[gapminder$continent == "Americas",]
ggplot(data = americas, mapping = aes(x = year, y = lifeExp)) +
geom_line() +
facet_wrap( ~ country) +
theme(axis.text.x = element_text(angle = 45))
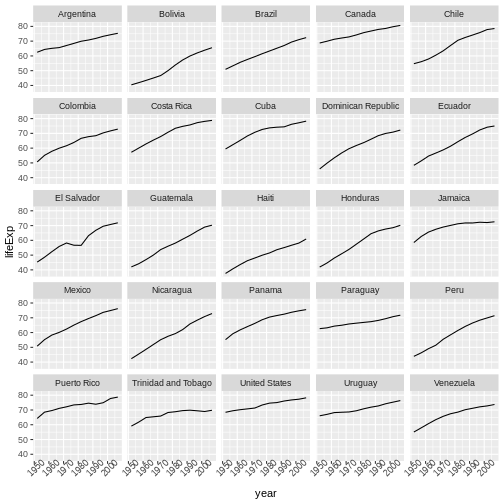
The facet_wrap layer took a “formula” as its argument,
denoted by the tilde (~). This tells R to draw a panel for each unique
value in the country column of the gapminder dataset.
Modifying text
To clean this figure up for a publication we need to change some of the text elements. The x-axis is too cluttered, and the y axis should read “Life expectancy”, rather than the column name in the data frame.
We can do this by adding a couple of different layers. The
theme layer controls the axis text, and overall text
size. Labels for the axes, plot title and any legend can be set using
the labs function. Legend titles are set using the same
names we used in the aes specification. Thus below the
color legend title is set using color = "Continent", while
the title of a fill legend would be set using
fill = "MyTitle".
R
ggplot(data = americas, mapping = aes(x = year, y = lifeExp, color=continent)) +
geom_line() + facet_wrap( ~ country) +
labs(
x = "Year", # x axis title
y = "Life expectancy", # y axis title
title = "Figure 1", # main title of figure
color = "Continent" # title of legend
) +
theme(axis.text.x = element_text(angle = 90, hjust = 1))
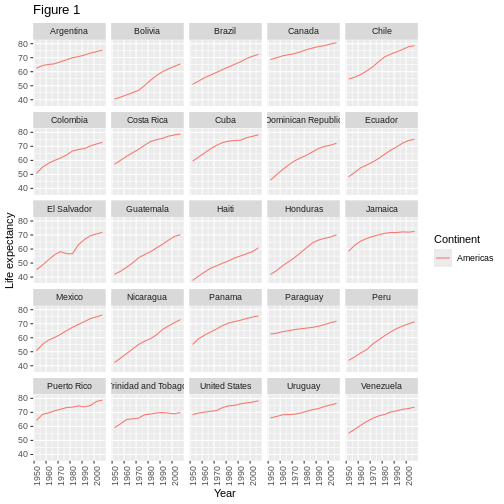
Exporting the plot
The ggsave() function allows you to export a plot
created with ggplot. You can specify the dimension and resolution of
your plot by adjusting the appropriate arguments (width,
height and dpi) to create high quality
graphics for publication. In order to save the plot from above, we first
assign it to a variable lifeExp_plot, then tell
ggsave to save that plot in png format to a
directory called results. (Make sure you have a
results/ folder in your working directory.)
R
lifeExp_plot <- ggplot(data = americas, mapping = aes(x = year, y = lifeExp, color=continent)) +
geom_line() + facet_wrap( ~ country) +
labs(
x = "Year", # x axis title
y = "Life expectancy", # y axis title
title = "Figure 1", # main title of figure
color = "Continent" # title of legend
) +
theme(axis.text.x = element_text(angle = 90, hjust = 1))
ggsave(filename = "results/lifeExp.png", plot = lifeExp_plot, width = 12, height = 10, dpi = 300, units = "cm")
There are two nice things about ggsave. First, it
defaults to the last plot, so if you omit the plot argument
it will automatically save the last plot you created with
ggplot. Secondly, it tries to determine the format you want
to save your plot in from the file extension you provide for the
filename (for example .png or .pdf). If you
need to, you can specify the format explicitly in the
device argument.
This is a taste of what you can do with ggplot2. RStudio provides a really useful cheat sheet of the different layers available, and more extensive documentation is available on the ggplot2 website. All RStudio cheat sheets are available from the RStudio website. Finally, if you have no idea how to change something, a quick Google search will usually send you to a relevant question and answer on Stack Overflow with reusable code to modify!
Challenge 5
Generate boxplots to compare life expectancy between the different continents during the available years.
Advanced:
- Rename y axis as Life Expectancy.
- Remove x axis labels.
Here a possible solution: xlab() and ylab()
set labels for the x and y axes, respectively The axis title, text and
ticks are attributes of the theme and must be modified within a
theme() call.
R
ggplot(data = gapminder, mapping = aes(x = continent, y = lifeExp, fill = continent)) +
geom_boxplot() + facet_wrap(~year) +
ylab("Life Expectancy") +
theme(axis.title.x=element_blank(),
axis.text.x = element_blank(),
axis.ticks.x = element_blank())
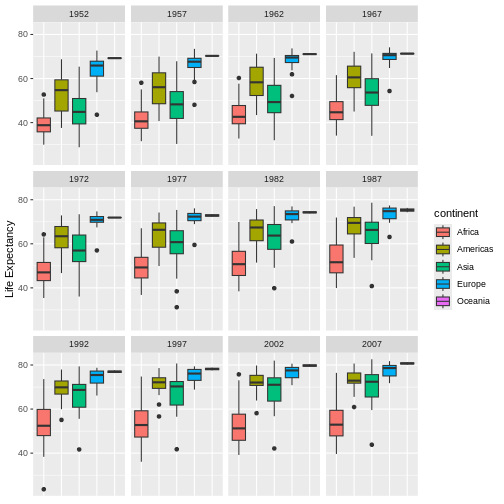
- Use
ggplot2to create plots. - Think about graphics in layers: aesthetics, geometry, statistics, scale transformation, and grouping.
Content from Vectorization
Last updated on 2026-04-21 | Edit this page
Estimated time: 25 minutes
Overview
Questions
- How can I operate on all the elements of a vector at once?
Objectives
- To understand vectorized operations in R.
Most of R’s functions are vectorized, meaning that the function will operate on all elements of a vector without needing to loop through and act on each element one at a time. This makes writing code more concise, easy to read, and less error prone.
R
x <- 1:4
x * 2
OUTPUT
[1] 2 4 6 8The multiplication happened to each element of the vector.
We can also add two vectors together:
R
y <- 6:9
x + y
OUTPUT
[1] 7 9 11 13Each element of x was added to its corresponding element
of y:
Here is how we would add two vectors together using a for loop:
R
output_vector <- c()
for (i in 1:4) {
output_vector[i] <- x[i] + y[i]
}
output_vector
OUTPUT
[1] 7 9 11 13Compare this to the output using vectorised operations.
R
sum_xy <- x + y
sum_xy
OUTPUT
[1] 7 9 11 13Challenge 1
Let’s try this on the pop column of the
gapminder dataset.
Make a new column in the gapminder data frame that
contains population in units of millions of people. Check the head or
tail of the data frame to make sure it worked.
Let’s try this on the pop column of the
gapminder dataset.
Make a new column in the gapminder data frame that
contains population in units of millions of people. Check the head or
tail of the data frame to make sure it worked.
R
gapminder$pop_millions <- gapminder$pop / 1e6
head(gapminder)
OUTPUT
country year pop continent lifeExp gdpPercap pop_millions
1 Afghanistan 1952 8425333 Asia 28.801 779.4453 8.425333
2 Afghanistan 1957 9240934 Asia 30.332 820.8530 9.240934
3 Afghanistan 1962 10267083 Asia 31.997 853.1007 10.267083
4 Afghanistan 1967 11537966 Asia 34.020 836.1971 11.537966
5 Afghanistan 1972 13079460 Asia 36.088 739.9811 13.079460
6 Afghanistan 1977 14880372 Asia 38.438 786.1134 14.880372Challenge 2
On a single graph, plot population, in millions, against year, for all countries. Do not worry about identifying which country is which.
Repeat the exercise, graphing only for China, India, and Indonesia. Again, do not worry about which is which.
Refresh your plotting skills by plotting population in millions against year.
R
ggplot(gapminder, aes(x = year, y = pop_millions)) +
geom_point()
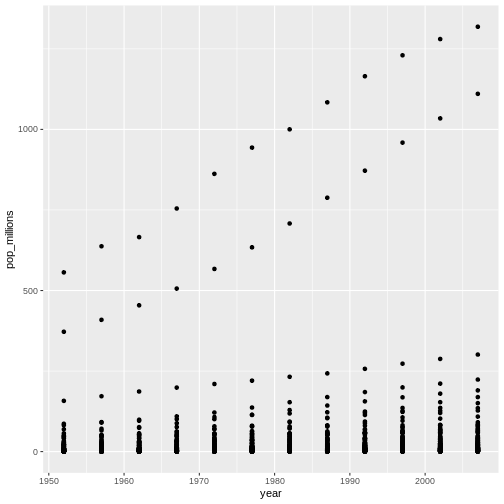
R
countryset <- c("China","India","Indonesia")
ggplot(gapminder[gapminder$country %in% countryset,],
aes(x = year, y = pop_millions)) +
geom_point()
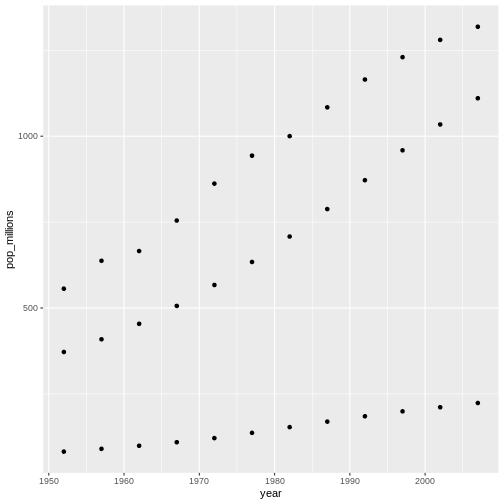
Comparison operators, logical operators, and many functions are also vectorized:
Comparison operators
R
x > 2
OUTPUT
[1] FALSE FALSE TRUE TRUELogical operators
R
a <- x > 3 # or, for clarity, a <- (x > 3)
a
OUTPUT
[1] FALSE FALSE FALSE TRUETip: some useful functions for logical vectors
any() will return TRUE if any
element of a vector is TRUE.all() will return TRUE if all
elements of a vector are TRUE.
Most functions also operate element-wise on vectors:
Functions
R
x <- 1:4
log(x)
OUTPUT
[1] 0.0000000 0.6931472 1.0986123 1.3862944Vectorized operations work element-wise on matrices:
R
m <- matrix(1:12, nrow=3, ncol=4)
m * -1
OUTPUT
[,1] [,2] [,3] [,4]
[1,] -1 -4 -7 -10
[2,] -2 -5 -8 -11
[3,] -3 -6 -9 -12Tip: element-wise vs. matrix multiplication
Very important: the operator * gives you element-wise
multiplication! To do matrix multiplication, we need to use the
%*% operator:
R
m %*% matrix(1, nrow=4, ncol=1)
OUTPUT
[,1]
[1,] 22
[2,] 26
[3,] 30R
matrix(1:4, nrow=1) %*% matrix(1:4, ncol=1)
OUTPUT
[,1]
[1,] 30For more on matrix algebra, see the Quick-R reference guide
Challenge 3
Given the following matrix:
R
m <- matrix(1:12, nrow=3, ncol=4)
m
OUTPUT
[,1] [,2] [,3] [,4]
[1,] 1 4 7 10
[2,] 2 5 8 11
[3,] 3 6 9 12Write down what you think will happen when you run:
m ^ -1m * c(1, 0, -1)m > c(0, 20)m * c(1, 0, -1, 2)
Did you get the output you expected? If not, ask a helper!
Given the following matrix:
R
m <- matrix(1:12, nrow=3, ncol=4)
m
OUTPUT
[,1] [,2] [,3] [,4]
[1,] 1 4 7 10
[2,] 2 5 8 11
[3,] 3 6 9 12Write down what you think will happen when you run:
m ^ -1
OUTPUT
[,1] [,2] [,3] [,4]
[1,] 1.0000000 0.2500000 0.1428571 0.10000000
[2,] 0.5000000 0.2000000 0.1250000 0.09090909
[3,] 0.3333333 0.1666667 0.1111111 0.08333333m * c(1, 0, -1)
OUTPUT
[,1] [,2] [,3] [,4]
[1,] 1 4 7 10
[2,] 0 0 0 0
[3,] -3 -6 -9 -12m > c(0, 20)
OUTPUT
[,1] [,2] [,3] [,4]
[1,] TRUE FALSE TRUE FALSE
[2,] FALSE TRUE FALSE TRUE
[3,] TRUE FALSE TRUE FALSEChallenge 4
We’re interested in looking at the sum of the following sequence of fractions:
R
x = 1/(1^2) + 1/(2^2) + 1/(3^2) + ... + 1/(n^2)
This would be tedious to type out, and impossible for high values of n. Use vectorisation to compute x when n=100. What is the sum when n=10,000?
We’re interested in looking at the sum of the following sequence of fractions:
R
x = 1/(1^2) + 1/(2^2) + 1/(3^2) + ... + 1/(n^2)
This would be tedious to type out, and impossible for high values of n. Can you use vectorisation to compute x, when n=100? How about when n=10,000?
R
sum(1/(1:100)^2)
OUTPUT
[1] 1.634984R
sum(1/(1:1e04)^2)
OUTPUT
[1] 1.644834R
n <- 10000
sum(1/(1:n)^2)
OUTPUT
[1] 1.644834We can also obtain the same results using a function:
R
inverse_sum_of_squares <- function(n) {
sum(1/(1:n)^2)
}
inverse_sum_of_squares(100)
OUTPUT
[1] 1.634984R
inverse_sum_of_squares(10000)
OUTPUT
[1] 1.644834R
n <- 10000
inverse_sum_of_squares(n)
OUTPUT
[1] 1.644834Tip: Operations on vectors of unequal length
Operations can also be performed on vectors of unequal length, through a process known as recycling. This process automatically repeats the smaller vector until it matches the length of the larger vector. R will provide a warning if the larger vector is not a multiple of the smaller vector.
R
x <- c(1, 2, 3)
y <- c(1, 2, 3, 4, 5, 6, 7)
x + y
WARNING
Warning in x + y: longer object length is not a multiple of shorter object
lengthOUTPUT
[1] 2 4 6 5 7 9 8Vector x was recycled to match the length of vector
y
- Use vectorized operations instead of loops.
Content from Functions Explained
Last updated on 2026-04-21 | Edit this page
Estimated time: 60 minutes
Overview
Questions
- How can I write a new function in R?
Objectives
- Define a function that takes arguments.
- Return a value from a function.
- Check argument conditions with
stopifnot()in functions. - Test a function.
- Set default values for function arguments.
- Explain why we should divide programs into small, single-purpose functions.
If we only had one data set to analyze, it would probably be faster to load the file into a spreadsheet and use that to plot simple statistics. However, the gapminder data is updated periodically, and we may want to pull in that new information later and re-run our analysis again. We may also obtain similar data from a different source in the future.
In this lesson, we’ll learn how to write a function so that we can repeat several operations with a single command.
What is a function?
Functions gather a sequence of operations into a whole, preserving it for ongoing use. Functions provide:
- a name we can remember and invoke it by
- relief from the need to remember the individual operations
- a defined set of inputs and expected outputs
- rich connections to the larger programming environment
As the basic building block of most programming languages, user-defined functions constitute “programming” as much as any single abstraction can. If you have written a function, you are a computer programmer.
Defining a function
Let’s open a new R script file in the functions/
directory and call it functions-lesson.R.
The general structure of a function is:
R
my_function <- function(parameters) {
# perform action
# return value
}
Let’s define a function fahr_to_kelvin() that converts
temperatures from Fahrenheit to Kelvin:
R
fahr_to_kelvin <- function(temp) {
kelvin <- ((temp - 32) * (5 / 9)) + 273.15
return(kelvin)
}
We define fahr_to_kelvin() by assigning it to the output
of function. The list of argument names are contained
within parentheses. Next, the body of
the function–the statements that are executed when it runs–is contained
within curly braces ({}). The statements in the body are
indented by two spaces. This makes the code easier to read but does not
affect how the code operates.
It is useful to think of creating functions like writing a cookbook. First you define the “ingredients” that your function needs. In this case, we only need one ingredient to use our function: “temp”. After we list our ingredients, we then say what we will do with them, in this case, we are taking our ingredient and applying a set of mathematical operators to it.
When we call the function, the values we pass to it as arguments are assigned to those variables so that we can use them inside the function. Inside the function, we use a return statement to send a result back to whoever asked for it.
Tip
One feature unique to R is that the return statement is not required. R automatically returns whichever variable is on the last line of the body of the function. But for clarity, we will explicitly define the return statement.
Let’s try running our function. Calling our own function is no different from calling any other function:
R
# freezing point of water
fahr_to_kelvin(32)
OUTPUT
[1] 273.15R
# boiling point of water
fahr_to_kelvin(212)
OUTPUT
[1] 373.15Challenge 1
Write a function called kelvin_to_celsius() that takes a
temperature in Kelvin and returns that temperature in Celsius.
Hint: To convert from Kelvin to Celsius you subtract 273.15
Write a function called kelvin_to_celsius that takes a
temperature in Kelvin and returns that temperature in Celsius
R
kelvin_to_celsius <- function(temp) {
celsius <- temp - 273.15
return(celsius)
}
Combining functions
The real power of functions comes from mixing, matching and combining them into ever-larger chunks to get the effect we want.
Let’s define two functions that will convert temperature from Fahrenheit to Kelvin, and Kelvin to Celsius:
R
fahr_to_kelvin <- function(temp) {
kelvin <- ((temp - 32) * (5 / 9)) + 273.15
return(kelvin)
}
kelvin_to_celsius <- function(temp) {
celsius <- temp - 273.15
return(celsius)
}
Challenge 2
Define the function to convert directly from Fahrenheit to Celsius, by reusing the two functions above (or using your own functions if you prefer).
Define the function to convert directly from Fahrenheit to Celsius, by reusing these two functions above
R
fahr_to_celsius <- function(temp) {
temp_k <- fahr_to_kelvin(temp)
result <- kelvin_to_celsius(temp_k)
return(result)
}
Interlude: Defensive Programming
Now that we’ve begun to appreciate how writing functions provides an
efficient way to make R code re-usable and modular, we should note that
it is important to ensure that functions only work in their intended
use-cases. Checking function parameters is related to the concept of
defensive programming. Defensive programming encourages us to
frequently check conditions and throw an error if something is wrong.
These checks are referred to as assertion statements because we want to
assert some condition is TRUE before proceeding. They make
it easier to debug because they give us a better idea of where the
errors originate.
Checking conditions with stopifnot()
Let’s start by re-examining fahr_to_kelvin(), our
function for converting temperatures from Fahrenheit to Kelvin. It was
defined like so:
R
fahr_to_kelvin <- function(temp) {
kelvin <- ((temp - 32) * (5 / 9)) + 273.15
return(kelvin)
}
For this function to work as intended, the argument temp
must be a numeric value; otherwise, the mathematical
procedure for converting between the two temperature scales will not
work. To create an error, we can use the function stop().
For example, since the argument temp must be a
numeric vector, we could check for this condition with an
if statement and throw an error if the condition was
violated. We could augment our function above like so:
R
fahr_to_kelvin <- function(temp) {
if (!is.numeric(temp)) {
stop("temp must be a numeric vector.")
}
kelvin <- ((temp - 32) * (5 / 9)) + 273.15
return(kelvin)
}
If we had multiple conditions or arguments to check, it would take
many lines of code to check all of them. Luckily R provides the
convenience function stopifnot(). We can list as many
requirements that should evaluate to TRUE;
stopifnot() throws an error if it finds one that is
FALSE. Listing these conditions also serves a secondary
purpose as extra documentation for the function.
Let’s try out defensive programming with stopifnot() by
adding assertions to check the input to our function
fahr_to_kelvin().
We want to assert the following: temp is a numeric
vector. We may do that like so:
R
fahr_to_kelvin <- function(temp) {
stopifnot(is.numeric(temp))
kelvin <- ((temp - 32) * (5 / 9)) + 273.15
return(kelvin)
}
It still works when given proper input.
R
# freezing point of water
fahr_to_kelvin(temp = 32)
OUTPUT
[1] 273.15But fails instantly if given improper input.
R
# Metric is a factor instead of numeric
fahr_to_kelvin(temp = as.factor(32))
ERROR
Error in `fahr_to_kelvin()`:
! is.numeric(temp) is not TRUEChallenge 3
Use defensive programming to ensure that our
fahr_to_celsius() function throws an error immediately if
the argument temp is specified inappropriately.
Extend our previous definition of the function by adding in an
explicit call to stopifnot(). Since
fahr_to_celsius() is a composition of two other functions,
checking inside here makes adding checks to the two component functions
redundant.
R
fahr_to_celsius <- function(temp) {
stopifnot(is.numeric(temp))
temp_k <- fahr_to_kelvin(temp)
result <- kelvin_to_celsius(temp_k)
return(result)
}
More on combining functions
Now, we’re going to define a function that calculates the Gross Domestic Product of a nation from the data available in our dataset:
R
# Takes a dataset and multiplies the population column
# with the GDP per capita column.
calcGDP <- function(dat) {
gdp <- dat$pop * dat$gdpPercap
return(gdp)
}
We define calcGDP() by assigning it to the output of
function. The list of argument names are contained within
parentheses. Next, the body of the function -- the statements executed
when you call the function – is contained within curly braces
({}).
We’ve indented the statements in the body by two spaces. This makes the code easier to read but does not affect how it operates.
When we call the function, the values we pass to it are assigned to the arguments, which become variables inside the body of the function.
Inside the function, we use the return() function to
send back the result. This return() function is optional: R
will automatically return the results of whatever command is executed on
the last line of the function.
R
calcGDP(head(gapminder))
OUTPUT
[1] 6567086330 7585448670 8758855797 9648014150 9678553274 11697659231That’s not very informative. Let’s add some more arguments so we can extract that per year and country.
R
# Takes a dataset and multiplies the population column
# with the GDP per capita column.
calcGDP <- function(dat, year=NULL, country=NULL) {
if(!is.null(year)) {
dat <- dat[dat$year %in% year, ]
}
if (!is.null(country)) {
dat <- dat[dat$country %in% country,]
}
gdp <- dat$pop * dat$gdpPercap
new <- cbind(dat, gdp=gdp)
return(new)
}
If you’ve been writing these functions down into a separate R script
(a good idea!), you can load in the functions into our R session by
using the source() function:
R
source("functions/functions-lesson.R")
Ok, so there’s a lot going on in this function now. In plain English, the function now subsets the provided data by year if the year argument isn’t empty, then subsets the result by country if the country argument isn’t empty. Then it calculates the GDP for whatever subset emerges from the previous two steps. The function then adds the GDP as a new column to the subsetted data and returns this as the final result. You can see that the output is much more informative than a vector of numbers.
Let’s take a look at what happens when we specify the year:
R
head(calcGDP(gapminder, year=2007))
OUTPUT
country year pop continent lifeExp gdpPercap gdp
12 Afghanistan 2007 31889923 Asia 43.828 974.5803 31079291949
24 Albania 2007 3600523 Europe 76.423 5937.0295 21376411360
36 Algeria 2007 33333216 Africa 72.301 6223.3675 207444851958
48 Angola 2007 12420476 Africa 42.731 4797.2313 59583895818
60 Argentina 2007 40301927 Americas 75.320 12779.3796 515033625357
72 Australia 2007 20434176 Oceania 81.235 34435.3674 703658358894Or for a specific country:
R
calcGDP(gapminder, country="Australia")
OUTPUT
country year pop continent lifeExp gdpPercap gdp
61 Australia 1952 8691212 Oceania 69.120 10039.60 87256254102
62 Australia 1957 9712569 Oceania 70.330 10949.65 106349227169
63 Australia 1962 10794968 Oceania 70.930 12217.23 131884573002
64 Australia 1967 11872264 Oceania 71.100 14526.12 172457986742
65 Australia 1972 13177000 Oceania 71.930 16788.63 221223770658
66 Australia 1977 14074100 Oceania 73.490 18334.20 258037329175
67 Australia 1982 15184200 Oceania 74.740 19477.01 295742804309
68 Australia 1987 16257249 Oceania 76.320 21888.89 355853119294
69 Australia 1992 17481977 Oceania 77.560 23424.77 409511234952
70 Australia 1997 18565243 Oceania 78.830 26997.94 501223252921
71 Australia 2002 19546792 Oceania 80.370 30687.75 599847158654
72 Australia 2007 20434176 Oceania 81.235 34435.37 703658358894Or both:
R
calcGDP(gapminder, year=2007, country="Australia")
OUTPUT
country year pop continent lifeExp gdpPercap gdp
72 Australia 2007 20434176 Oceania 81.235 34435.37 703658358894Let’s walk through the body of the function:
Here we’ve added two arguments, year, and
country. We’ve set default arguments for both as
NULL using the = operator in the function
definition. This means that those arguments will take on those values
unless the user specifies otherwise.
R
if(!is.null(year)) {
dat <- dat[dat$year %in% year, ]
}
if (!is.null(country)) {
dat <- dat[dat$country %in% country,]
}
Here, we check whether each additional argument is set to
null, and whenever they’re not null overwrite
the dataset stored in dat with a subset given by the
non-null argument.
Building these conditionals into the function makes it more flexible for later. Now, we can use it to calculate the GDP for:
- The whole dataset;
- A single year;
- A single country;
- A single combination of year and country.
By using %in% instead, we can also give multiple years
or countries to those arguments.
Tip: Pass by value
Functions in R almost always make copies of the data to operate on
inside of a function body. When we modify dat inside the
function we are modifying the copy of the gapminder dataset stored in
dat, not the original variable we gave as the first
argument.
This is called “pass-by-value” and it makes writing code much safer: you can always be sure that whatever changes you make within the body of the function, stay inside the body of the function.
Tip: Function scope
Another important concept is scoping: any variables (or functions!)
you create or modify inside the body of a function only exist for the
lifetime of the function’s execution. When we call
calcGDP(), the variables dat, gdp
and new only exist inside the body of the function. Even if
we have variables of the same name in our interactive R session, they
are not modified in any way when executing a function.
Finally, we calculated the GDP on our new subset, and created a new data frame with that column added. This means when we call the function later we can see the context for the returned GDP values, which is much better than in our first attempt where we got a vector of numbers.
Challenge 4
Test out your GDP function by calculating the GDP for New Zealand in 1987. How does this differ from New Zealand’s GDP in 1952?
R
calcGDP(gapminder, year = c(1952, 1987), country = "New Zealand")
GDP for New Zealand in 1987: 65050008703
GDP for New Zealand in 1952: 21058193787
Challenge 5
The paste() function can be used to combine text
together, e.g:
R
best_practice <- c("Write", "programs", "for", "people", "not", "computers")
paste(best_practice, collapse=" ")
OUTPUT
[1] "Write programs for people not computers"Write a function called fence() that takes two vectors
as arguments, called text and wrapper, and
prints out the text wrapped with the wrapper:
R
fence(text=best_practice, wrapper="***")
Note: the paste() function has an argument
called sep, which specifies the separator between text. The
default is a space: ” “. The default for paste0() is no
space”“.
Write a function called fence() that takes two vectors
as arguments, called text and wrapper, and
prints out the text wrapped with the wrapper:
R
fence <- function(text, wrapper){
text <- c(wrapper, text, wrapper)
result <- paste(text, collapse = " ")
return(result)
}
best_practice <- c("Write", "programs", "for", "people", "not", "computers")
fence(text=best_practice, wrapper="***")
OUTPUT
[1] "*** Write programs for people not computers ***"Tip
R has some unique aspects that can be exploited when performing more complicated operations. We will not be writing anything that requires knowledge of these more advanced concepts. In the future when you are comfortable writing functions in R, you can learn more by reading the R Language Manual or this chapter from Advanced R Programming by Hadley Wickham.
Tip: Testing and documenting
It’s important to both test functions and document them: Documentation helps you, and others, understand what the purpose of your function is, and how to use it, and its important to make sure that your function actually does what you think.
When you first start out, your workflow will probably look a lot like this:
- Write a function
- Comment parts of the function to document its behaviour
- Load in the source file
- Experiment with it in the console to make sure it behaves as you expect
- Make any necessary bug fixes
- Rinse and repeat.
Formal documentation for functions, written in separate
.Rd files, gets turned into the documentation you see in
help files. The roxygen2
package allows R coders to write documentation alongside the function
code and then process it into the appropriate .Rd files.
You will want to switch to this more formal method of writing
documentation when you start writing more complicated R projects. In
fact, packages are, in essence, bundles of functions with this formal
documentation. Loading your own functions through
source("functions.R") is equivalent to loading someone
else’s functions (or your own one day!) through
library("package").
Formal automated tests can be written using the testthat package.
- Use
functionto define a new function in R. - Use parameters to pass values into functions.
- Use
stopifnot()to flexibly check function arguments in R. - Load functions into programs using
source().
Content from Writing Data
Last updated on 2026-04-21 | Edit this page
Estimated time: 20 minutes
Overview
Questions
- How can I save plots and data created in R?
Objectives
- To be able to write out plots and data from R.
Saving plots
You have already seen how to save the most recent plot you create in
ggplot2, using the command ggsave. As a
refresher:
R
ggsave("My_most_recent_plot.pdf")
You can save a plot from within RStudio using the ‘Export’ button in the ‘Plot’ window. This will give you the option of saving as a .pdf or as .png, .jpg or other image formats.
Sometimes you will want to save plots without creating them in the ‘Plot’ window first. Perhaps you want to make a pdf document with multiple pages: each one a different plot, for example. Or perhaps you’re looping through multiple subsets of a file, plotting data from each subset, and you want to save each plot, but obviously can’t stop the loop to click ‘Export’ for each one.
In this case you can use a more flexible approach. The function
pdf creates a new pdf device. You can control the size and
resolution using the arguments to this function.
R
pdf("Life_Exp_vs_time.pdf", width=12, height=4)
ggplot(data=gapminder, aes(x=year, y=lifeExp, colour=country)) +
geom_line() +
theme(legend.position = "none")
# You then have to make sure to turn off the pdf device!
dev.off()
Open up this document and have a look.
Challenge 1
Rewrite your ‘pdf’ command to print a second page in the pdf, showing
a facet plot (hint: use facet_grid) of the same data with
one panel per continent.
R
pdf("Life_Exp_vs_time.pdf", width = 12, height = 4)
p <- ggplot(data = gapminder, aes(x = year, y = lifeExp, colour = country)) +
geom_line() +
theme(legend.position = "none")
p
p + facet_grid(~continent)
dev.off()
The commands jpeg, png etc. are used
similarly to produce documents in different formats.
Writing data
At some point, you’ll also want to write out data from R.
We can use the write.table function for this, which is
very similar to read.table from before.
Let’s create a data-cleaning script, for this analysis, we only want to focus on the gapminder data for Australia:
R
aust_subset <- gapminder[gapminder$country == "Australia",]
write.table(aust_subset,
file="cleaned-data/gapminder-aus.csv",
sep=","
)
Let’s switch back to the shell to take a look at the data to make sure it looks OK:
OUTPUT
"country","year","pop","continent","lifeExp","gdpPercap"
"61","Australia",1952,8691212,"Oceania",69.12,10039.59564
"62","Australia",1957,9712569,"Oceania",70.33,10949.64959
"63","Australia",1962,10794968,"Oceania",70.93,12217.22686
"64","Australia",1967,11872264,"Oceania",71.1,14526.12465
"65","Australia",1972,13177000,"Oceania",71.93,16788.62948
"66","Australia",1977,14074100,"Oceania",73.49,18334.19751
"67","Australia",1982,15184200,"Oceania",74.74,19477.00928
"68","Australia",1987,16257249,"Oceania",76.32,21888.88903
"69","Australia",1992,17481977,"Oceania",77.56,23424.76683Hmm, that’s not quite what we wanted. Where did all these quotation marks come from? Also the row numbers are meaningless.
Let’s look at the help file to work out how to change this behaviour.
R
?write.table
By default R will wrap character vectors with quotation marks when writing out to file. It will also write out the row and column names.
Let’s fix this:
R
write.table(
gapminder[gapminder$country == "Australia",],
file="cleaned-data/gapminder-aus.csv",
sep=",", quote=FALSE, row.names=FALSE
)
Now lets look at the data again using our shell skills:
OUTPUT
country,year,pop,continent,lifeExp,gdpPercap
Australia,1952,8691212,Oceania,69.12,10039.59564
Australia,1957,9712569,Oceania,70.33,10949.64959
Australia,1962,10794968,Oceania,70.93,12217.22686
Australia,1967,11872264,Oceania,71.1,14526.12465
Australia,1972,13177000,Oceania,71.93,16788.62948
Australia,1977,14074100,Oceania,73.49,18334.19751
Australia,1982,15184200,Oceania,74.74,19477.00928
Australia,1987,16257249,Oceania,76.32,21888.88903
Australia,1992,17481977,Oceania,77.56,23424.76683That looks better!
Challenge 2
Write a data-cleaning script file that subsets the gapminder data to include only data points collected since 1990.
Use this script to write out the new subset to a file in the
cleaned-data/ directory.
R
write.table(
gapminder[gapminder$year > 1990, ],
file = "cleaned-data/gapminder-after1990.csv",
sep = ",", quote = FALSE, row.names = FALSE
)
- Save plots from RStudio using the ‘Export’ button.
- Use
write.tableto save tabular data.
Content from Data Frame Manipulation with dplyr
Last updated on 2026-04-21 | Edit this page
Estimated time: 55 minutes
Overview
Questions
- How can I manipulate data frames without repeating myself?
Objectives
- To be able to use the six main data frame manipulation ‘verbs’ with
pipes in
dplyr. - To understand how
group_by()andsummarize()can be combined to summarize datasets. - Be able to analyze a subset of data using logical filtering.
Manipulation of data frames means many things to many researchers: we often select certain observations (rows) or variables (columns), we often group the data by a certain variable(s), or we even calculate summary statistics. We can do these operations using the normal base R operations:
R
mean(gapminder$gdpPercap[gapminder$continent == "Africa"])
OUTPUT
[1] 2193.755R
mean(gapminder$gdpPercap[gapminder$continent == "Americas"])
OUTPUT
[1] 7136.11R
mean(gapminder$gdpPercap[gapminder$continent == "Asia"])
OUTPUT
[1] 7902.15But this isn’t very nice because there is a fair bit of repetition. Repeating yourself will cost you time, both now and later, and potentially introduce some nasty bugs.
The dplyr package
Luckily, the dplyr
package provides a number of very useful functions for manipulating data
frames in a way that will reduce the above repetition, reduce the
probability of making errors, and probably even save you some typing. As
an added bonus, you might even find the dplyr grammar
easier to read.
Tip: Tidyverse
dplyr package belongs to a broader family of opinionated
R packages designed for data science called the “Tidyverse”. These
packages are specifically designed to work harmoniously together. Some
of these packages will be covered along this course, but you can find
more complete information here: https://www.tidyverse.org/.
Here we’re going to cover 5 of the most commonly used functions as
well as using pipes (%>%) to combine them.
select()filter()group_by()summarize()mutate()
If you have have not installed this package earlier, please do so:
R
install.packages('dplyr')
Now let’s load the package:
R
library("dplyr")
Using select()
If, for example, we wanted to move forward with only a few of the
variables in our data frame we could use the select()
function. This will keep only the variables you select.
R
year_country_gdp <- select(gapminder, year, country, gdpPercap)
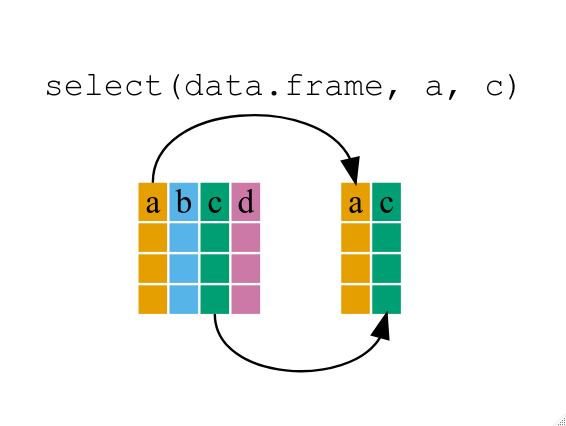 If we want to remove one column only from the
If we want to remove one column only from the gapminder
data, for example, removing the continent column.
R
smaller_gapminder_data <- select(gapminder, -continent)
If we open up year_country_gdp we’ll see that it only
contains the year, country and gdpPercap. Above we used ‘normal’
grammar, but the strengths of dplyr lie in combining
several functions using pipes. Since the pipes grammar is unlike
anything we’ve seen in R before, let’s repeat what we’ve done above
using pipes.
R
year_country_gdp <- gapminder %>% select(year, country, gdpPercap)
To help you understand why we wrote that in that way, let’s walk
through it step by step. First we summon the gapminder data frame and
pass it on, using the pipe symbol %>%, to the next step,
which is the select() function. In this case we don’t
specify which data object we use in the select() function
since in gets that from the previous pipe. Fun Fact:
There is a good chance you have encountered pipes before in the shell.
In R, a pipe symbol is %>% while in the shell it is
| but the concept is the same!
Tip: Renaming data frame columns in dplyr
In Chapter 4 we covered how you can rename columns with base R by
assigning a value to the output of the names() function.
Just like select, this is a bit cumbersome, but thankfully dplyr has a
rename() function.
Within a pipeline, the syntax is
rename(new_name = old_name). For example, we may want to
rename the gdpPercap column name from our select()
statement above.
R
tidy_gdp <- year_country_gdp %>% rename(gdp_per_capita = gdpPercap)
head(tidy_gdp)
OUTPUT
year country gdp_per_capita
1 1952 Afghanistan 779.4453
2 1957 Afghanistan 820.8530
3 1962 Afghanistan 853.1007
4 1967 Afghanistan 836.1971
5 1972 Afghanistan 739.9811
6 1977 Afghanistan 786.1134Using filter()
If we now want to move forward with the above, but only with European
countries, we can combine select and
filter
R
year_country_gdp_euro <- gapminder %>%
filter(continent == "Europe") %>%
select(year, country, gdpPercap)
If we now want to show life expectancy of European countries but only for a specific year (e.g., 2007), we can do as below.
R
europe_lifeExp_2007 <- gapminder %>%
filter(continent == "Europe", year == 2007) %>%
select(country, lifeExp)
Challenge 1
Write a single command (which can span multiple lines and includes
pipes) that will produce a data frame that has the African values for
lifeExp, country and year, but
not for other Continents. How many rows does your data frame have and
why?
R
year_country_lifeExp_Africa <- gapminder %>%
filter(continent == "Africa") %>%
select(year, country, lifeExp)
As with last time, first we pass the gapminder data frame to the
filter() function, then we pass the filtered version of the
gapminder data frame to the select() function.
Note: The order of operations is very important in this
case. If we used ‘select’ first, filter would not be able to find the
variable continent since we would have removed it in the previous
step.
Using group_by()
Now, we were supposed to be reducing the error prone repetitiveness
of what can be done with base R, but up to now we haven’t done that
since we would have to repeat the above for each continent. Instead of
filter(), which will only pass observations that meet your
criteria (in the above: continent=="Europe"), we can use
group_by(), which will essentially use every unique
criteria that you could have used in filter.
R
str(gapminder)
OUTPUT
'data.frame': 1704 obs. of 6 variables:
$ country : chr "Afghanistan" "Afghanistan" "Afghanistan" "Afghanistan" ...
$ year : int 1952 1957 1962 1967 1972 1977 1982 1987 1992 1997 ...
$ pop : num 8425333 9240934 10267083 11537966 13079460 ...
$ continent: chr "Asia" "Asia" "Asia" "Asia" ...
$ lifeExp : num 28.8 30.3 32 34 36.1 ...
$ gdpPercap: num 779 821 853 836 740 ...R
str(gapminder %>% group_by(continent))
OUTPUT
gropd_df [1,704 × 6] (S3: grouped_df/tbl_df/tbl/data.frame)
$ country : chr [1:1704] "Afghanistan" "Afghanistan" "Afghanistan" "Afghanistan" ...
$ year : int [1:1704] 1952 1957 1962 1967 1972 1977 1982 1987 1992 1997 ...
$ pop : num [1:1704] 8425333 9240934 10267083 11537966 13079460 ...
$ continent: chr [1:1704] "Asia" "Asia" "Asia" "Asia" ...
$ lifeExp : num [1:1704] 28.8 30.3 32 34 36.1 ...
$ gdpPercap: num [1:1704] 779 821 853 836 740 ...
- attr(*, "groups")= tibble [5 × 2] (S3: tbl_df/tbl/data.frame)
..$ continent: chr [1:5] "Africa" "Americas" "Asia" "Europe" ...
..$ .rows : list<int> [1:5]
.. ..$ : int [1:624] 25 26 27 28 29 30 31 32 33 34 ...
.. ..$ : int [1:300] 49 50 51 52 53 54 55 56 57 58 ...
.. ..$ : int [1:396] 1 2 3 4 5 6 7 8 9 10 ...
.. ..$ : int [1:360] 13 14 15 16 17 18 19 20 21 22 ...
.. ..$ : int [1:24] 61 62 63 64 65 66 67 68 69 70 ...
.. ..@ ptype: int(0)
..- attr(*, ".drop")= logi TRUEYou will notice that the structure of the data frame where we used
group_by() (grouped_df) is not the same as the
original gapminder (data.frame). A
grouped_df can be thought of as a list where
each item in the listis a data.frame which
contains only the rows that correspond to the a particular value
continent (at least in the example above).
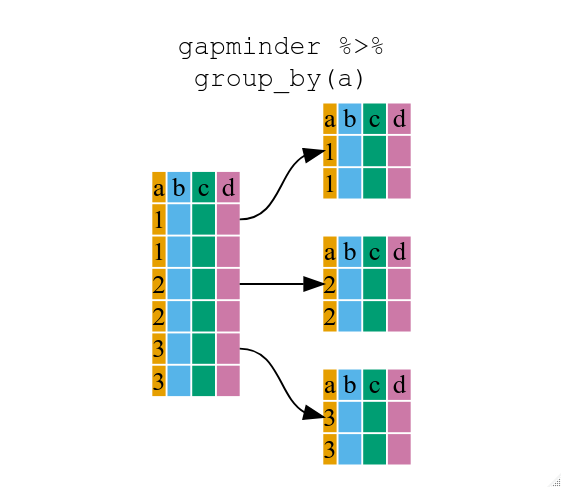
Using summarize()
The above was a bit on the uneventful side but
group_by() is much more exciting in conjunction with
summarize(). This will allow us to create new variable(s)
by using functions that repeat for each of the continent-specific data
frames. That is to say, using the group_by() function, we
split our original data frame into multiple pieces, then we can run
functions (e.g. mean() or sd()) within
summarize().
R
gdp_bycontinents <- gapminder %>%
group_by(continent) %>%
summarize(mean_gdpPercap = mean(gdpPercap))
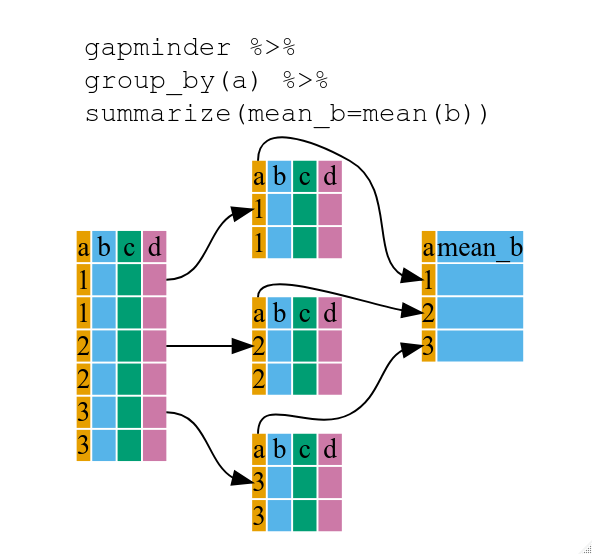
R
continent mean_gdpPercap
<fctr> <dbl>
1 Africa 2193.755
2 Americas 7136.110
3 Asia 7902.150
4 Europe 14469.476
5 Oceania 18621.609That allowed us to calculate the mean gdpPercap for each continent, but it gets even better.
Challenge 2
Calculate the average life expectancy per country. Which has the longest average life expectancy and which has the shortest average life expectancy?
R
lifeExp_bycountry <- gapminder %>%
group_by(country) %>%
summarize(mean_lifeExp = mean(lifeExp))
lifeExp_bycountry %>%
filter(mean_lifeExp == min(mean_lifeExp) | mean_lifeExp == max(mean_lifeExp))
OUTPUT
# A tibble: 2 × 2
country mean_lifeExp
<chr> <dbl>
1 Iceland 76.5
2 Sierra Leone 36.8Another way to do this is to use the dplyr function
arrange(), which arranges the rows in a data frame
according to the order of one or more variables from the data frame. It
has similar syntax to other functions from the dplyr
package. You can use desc() inside arrange()
to sort in descending order.
R
lifeExp_bycountry %>%
arrange(mean_lifeExp) %>%
head(1)
OUTPUT
# A tibble: 1 × 2
country mean_lifeExp
<chr> <dbl>
1 Sierra Leone 36.8R
lifeExp_bycountry %>%
arrange(desc(mean_lifeExp)) %>%
head(1)
OUTPUT
# A tibble: 1 × 2
country mean_lifeExp
<chr> <dbl>
1 Iceland 76.5Alphabetical order works too
R
lifeExp_bycountry %>%
arrange(desc(country)) %>%
head(1)
OUTPUT
# A tibble: 1 × 2
country mean_lifeExp
<chr> <dbl>
1 Zimbabwe 52.7The function group_by() allows us to group by multiple
variables. Let’s group by year and
continent.
R
gdp_bycontinents_byyear <- gapminder %>%
group_by(continent, year) %>%
summarize(mean_gdpPercap = mean(gdpPercap))
OUTPUT
`summarise()` has regrouped the output.
ℹ Summaries were computed grouped by continent and year.
ℹ Output is grouped by continent.
ℹ Use `summarise(.groups = "drop_last")` to silence this message.
ℹ Use `summarise(.by = c(continent, year))` for per-operation grouping
(`?dplyr::dplyr_by`) instead.That is already quite powerful, but it gets even better! You’re not
limited to defining 1 new variable in summarize().
R
gdp_pop_bycontinents_byyear <- gapminder %>%
group_by(continent, year) %>%
summarize(mean_gdpPercap = mean(gdpPercap),
sd_gdpPercap = sd(gdpPercap),
mean_pop = mean(pop),
sd_pop = sd(pop))
OUTPUT
`summarise()` has regrouped the output.
ℹ Summaries were computed grouped by continent and year.
ℹ Output is grouped by continent.
ℹ Use `summarise(.groups = "drop_last")` to silence this message.
ℹ Use `summarise(.by = c(continent, year))` for per-operation grouping
(`?dplyr::dplyr_by`) instead.count() and n()
A very common operation is to count the number of observations for
each group. The dplyr package comes with two related
functions that help with this.
For instance, if we wanted to check the number of countries included
in the dataset for the year 2002, we can use the count()
function. It takes the name of one or more columns that contain the
groups we are interested in, and we can optionally sort the results in
descending order by adding sort=TRUE:
R
gapminder %>%
filter(year == 2002) %>%
count(continent, sort = TRUE)
OUTPUT
continent n
1 Africa 52
2 Asia 33
3 Europe 30
4 Americas 25
5 Oceania 2If we need to use the number of observations in calculations, the
n() function is useful. It will return the total number of
observations in the current group rather than counting the number of
observations in each group within a specific column. For instance, if we
wanted to get the standard error of the life expectency per
continent:
R
gapminder %>%
group_by(continent) %>%
summarize(se_le = sd(lifeExp)/sqrt(n()))
OUTPUT
# A tibble: 5 × 2
continent se_le
<chr> <dbl>
1 Africa 0.366
2 Americas 0.540
3 Asia 0.596
4 Europe 0.286
5 Oceania 0.775You can also chain together several summary operations; in this case
calculating the minimum, maximum,
mean and se of each continent’s per-country
life-expectancy:
R
gapminder %>%
group_by(continent) %>%
summarize(
mean_le = mean(lifeExp),
min_le = min(lifeExp),
max_le = max(lifeExp),
se_le = sd(lifeExp)/sqrt(n()))
OUTPUT
# A tibble: 5 × 5
continent mean_le min_le max_le se_le
<chr> <dbl> <dbl> <dbl> <dbl>
1 Africa 48.9 23.6 76.4 0.366
2 Americas 64.7 37.6 80.7 0.540
3 Asia 60.1 28.8 82.6 0.596
4 Europe 71.9 43.6 81.8 0.286
5 Oceania 74.3 69.1 81.2 0.775Using mutate()
We can also create new variables prior to (or even after) summarizing
information using mutate().
R
gdp_pop_bycontinents_byyear <- gapminder %>%
mutate(gdp_billion = gdpPercap*pop/10^9) %>%
group_by(continent,year) %>%
summarize(mean_gdpPercap = mean(gdpPercap),
sd_gdpPercap = sd(gdpPercap),
mean_pop = mean(pop),
sd_pop = sd(pop),
mean_gdp_billion = mean(gdp_billion),
sd_gdp_billion = sd(gdp_billion))
OUTPUT
`summarise()` has regrouped the output.
ℹ Summaries were computed grouped by continent and year.
ℹ Output is grouped by continent.
ℹ Use `summarise(.groups = "drop_last")` to silence this message.
ℹ Use `summarise(.by = c(continent, year))` for per-operation grouping
(`?dplyr::dplyr_by`) instead.Connect mutate with logical filtering: ifelse
When creating new variables, we can hook this with a logical
condition. A simple combination of mutate() and
ifelse() facilitates filtering right where it is needed: in
the moment of creating something new. This easy-to-read statement is a
fast and powerful way of discarding certain data (even though the
overall dimension of the data frame will not change) or for updating
values depending on this given condition.
R
## keeping all data but "filtering" after a certain condition
# calculate GDP only for people with a life expectation above 25
gdp_pop_bycontinents_byyear_above25 <- gapminder %>%
mutate(gdp_billion = ifelse(lifeExp > 25, gdpPercap * pop / 10^9, NA)) %>%
group_by(continent, year) %>%
summarize(mean_gdpPercap = mean(gdpPercap),
sd_gdpPercap = sd(gdpPercap),
mean_pop = mean(pop),
sd_pop = sd(pop),
mean_gdp_billion = mean(gdp_billion),
sd_gdp_billion = sd(gdp_billion))
OUTPUT
`summarise()` has regrouped the output.
ℹ Summaries were computed grouped by continent and year.
ℹ Output is grouped by continent.
ℹ Use `summarise(.groups = "drop_last")` to silence this message.
ℹ Use `summarise(.by = c(continent, year))` for per-operation grouping
(`?dplyr::dplyr_by`) instead.R
## updating only if certain condition is fullfilled
# for life expectations above 40 years, the gpd to be expected in the future is scaled
gdp_future_bycontinents_byyear_high_lifeExp <- gapminder %>%
mutate(gdp_futureExpectation = ifelse(lifeExp > 40, gdpPercap * 1.5, gdpPercap)) %>%
group_by(continent, year) %>%
summarize(mean_gdpPercap = mean(gdpPercap),
mean_gdpPercap_expected = mean(gdp_futureExpectation))
OUTPUT
`summarise()` has regrouped the output.
ℹ Summaries were computed grouped by continent and year.
ℹ Output is grouped by continent.
ℹ Use `summarise(.groups = "drop_last")` to silence this message.
ℹ Use `summarise(.by = c(continent, year))` for per-operation grouping
(`?dplyr::dplyr_by`) instead.Combining dplyr and ggplot2
First install and load ggplot2:
R
install.packages('ggplot2')
R
library("ggplot2")
In the plotting lesson we looked at how to make a multi-panel figure
by adding a layer of facet panels using ggplot2. Here is
the code we used (with some extra comments):
R
# Filter countries located in the Americas
americas <- gapminder[gapminder$continent == "Americas", ]
# Make the plot
ggplot(data = americas, mapping = aes(x = year, y = lifeExp)) +
geom_line() +
facet_wrap( ~ country) +
theme(axis.text.x = element_text(angle = 45))
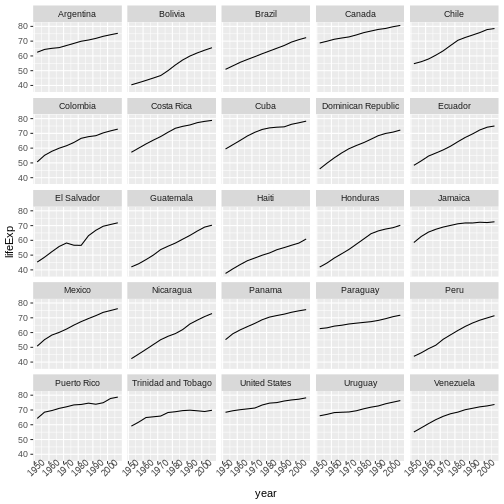
This code makes the right plot but it also creates an intermediate
variable (americas) that we might not have any other uses
for. Just as we used %>% to pipe data along a chain of
dplyr functions we can use it to pass data to
ggplot(). Because %>% replaces the first
argument in a function we don’t need to specify the data =
argument in the ggplot() function. By combining
dplyr and ggplot2 functions we can make the
same figure without creating any new variables or modifying the
data.
R
gapminder %>%
# Filter countries located in the Americas
filter(continent == "Americas") %>%
# Make the plot
ggplot(mapping = aes(x = year, y = lifeExp)) +
geom_line() +
facet_wrap( ~ country) +
theme(axis.text.x = element_text(angle = 45))
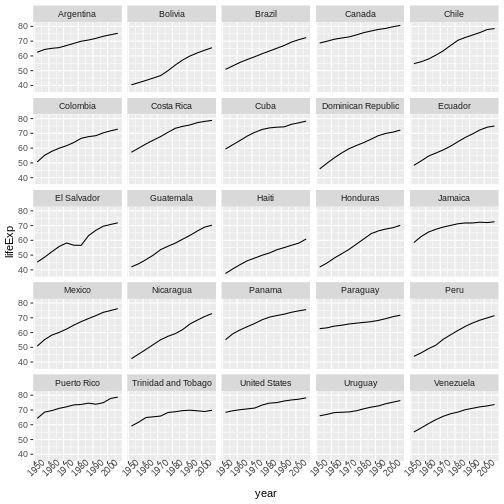
More examples of using the function mutate() and the
ggplot2 package.
R
gapminder %>%
# extract first letter of country name into new column
mutate(startsWith = substr(country, 1, 1)) %>%
# only keep countries starting with A or Z
filter(startsWith %in% c("A", "Z")) %>%
# plot lifeExp into facets
ggplot(aes(x = year, y = lifeExp, colour = continent)) +
geom_line() +
facet_wrap(vars(country)) +
theme_minimal()
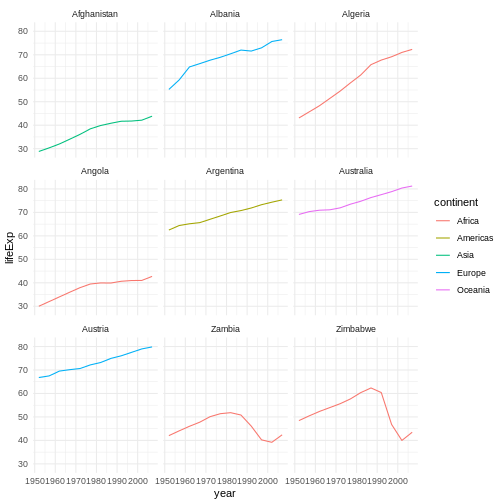
Advanced Challenge
Calculate the average life expectancy in 2002 of 2 randomly selected
countries for each continent. Then arrange the continent names in
reverse order. Hint: Use the dplyr
functions arrange() and slice_sample(), they
have similar syntax to other dplyr functions.
R
lifeExp_2countries_bycontinents <- gapminder %>%
filter(year==2002) %>%
group_by(continent) %>%
slice_sample(n=2) %>%
summarize(mean_lifeExp=mean(lifeExp)) %>%
arrange(desc(mean_lifeExp))
Other great resources
- R for Data Science (online book)
- Data Wrangling Cheat sheet (pdf file)
- Introduction to dplyr (online documentation)
- Data wrangling with R and RStudio (online video)
- Tidyverse Skills for Data Science (online book)
- Use the
dplyrpackage to manipulate data frames. - Use
select()to choose variables from a data frame. - Use
filter()to choose data based on values. - Use
group_by()andsummarize()to work with subsets of data. - Use
mutate()to create new variables.
Content from Data Frame Manipulation with tidyr
Last updated on 2026-04-21 | Edit this page
Estimated time: 45 minutes
Overview
Questions
- How can I change the layout of a data frame?
Objectives
- To understand the concepts of ‘longer’ and ‘wider’ data frame
formats and be able to convert between them with
tidyr.
Researchers often want to reshape their data frames from ‘wide’ to ‘longer’ layouts, or vice-versa. The ‘long’ layout or format is where:
- each column is a variable
- each row is an observation
In the purely ‘long’ (or ‘longest’) format, you usually have 1 column for the observed variable and the other columns are ID variables.
For the ‘wide’ format each row is often a site/subject/patient and
you have multiple observation variables containing the same type of
data. These can be either repeated observations over time, or
observation of multiple variables (or a mix of both). You may find data
input may be simpler or some other applications may prefer the ‘wide’
format. However, many of R‘s functions have been designed
assuming you have ’longer’ formatted data. This tutorial will help you
efficiently transform your data shape regardless of original format.
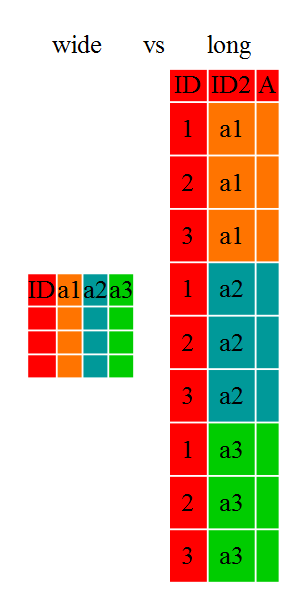
Long and wide data frame layouts mainly affect readability. For humans, the wide format is often more intuitive since we can often see more of the data on the screen due to its shape. However, the long format is more machine readable and is closer to the formatting of databases. The ID variables in our data frames are similar to the fields in a database and observed variables are like the database values.
Getting started
First install the packages if you haven’t already done so (you probably installed dplyr in the previous lesson):
R
#install.packages("tidyr")
#install.packages("dplyr")
Load the packages
R
library("tidyr")
library("dplyr")
First, lets look at the structure of our original gapminder data frame:
R
str(gapminder)
OUTPUT
'data.frame': 1704 obs. of 6 variables:
$ country : chr "Afghanistan" "Afghanistan" "Afghanistan" "Afghanistan" ...
$ year : int 1952 1957 1962 1967 1972 1977 1982 1987 1992 1997 ...
$ pop : num 8425333 9240934 10267083 11537966 13079460 ...
$ continent: chr "Asia" "Asia" "Asia" "Asia" ...
$ lifeExp : num 28.8 30.3 32 34 36.1 ...
$ gdpPercap: num 779 821 853 836 740 ...Challenge 1
Is gapminder a purely long, purely wide, or some intermediate format?
The original gapminder data.frame is in an intermediate format. It is
not purely long since it had multiple observation variables
(pop,lifeExp,gdpPercap).
Sometimes, as with the gapminder dataset, we have multiple types of
observed data. It is somewhere in between the purely ‘long’ and ‘wide’
data formats. We have 3 “ID variables” (continent,
country, year) and 3 “Observation variables”
(pop,lifeExp,gdpPercap). This
intermediate format can be preferred despite not having ALL observations
in 1 column given that all 3 observation variables have different units.
There are few operations that would need us to make this data frame any
longer (i.e. 4 ID variables and 1 Observation variable).
While using many of the functions in R, which are often vector based,
you usually do not want to do mathematical operations on values with
different units. For example, using the purely long format, a single
mean for all of the values of population, life expectancy, and GDP would
not be meaningful since it would return the mean of values with 3
incompatible units. The solution is that we first manipulate the data
either by grouping (see the lesson on dplyr), or we change
the structure of the data frame. Note: Some plotting
functions in R actually work better in the wide format data.
From wide to long format with pivot_longer()
Until now, we’ve been using the nicely formatted original gapminder dataset, but ‘real’ data (i.e. our own research data) will never be so well organized. Here let’s start with the wide formatted version of the gapminder dataset.
Download the wide version of the gapminder data from this link to a csv file and save it in your data folder.
We’ll load the data file and look at it. Note: we don’t want our
continent and country columns to be factors, so we use the
stringsAsFactors argument for read.csv() to disable
that.
R
gap_wide <- read.csv("data/gapminder_wide.csv", stringsAsFactors = FALSE)
str(gap_wide)
OUTPUT
'data.frame': 142 obs. of 38 variables:
$ continent : chr "Africa" "Africa" "Africa" "Africa" ...
$ country : chr "Algeria" "Angola" "Benin" "Botswana" ...
$ gdpPercap_1952: num 2449 3521 1063 851 543 ...
$ gdpPercap_1957: num 3014 3828 960 918 617 ...
$ gdpPercap_1962: num 2551 4269 949 984 723 ...
$ gdpPercap_1967: num 3247 5523 1036 1215 795 ...
$ gdpPercap_1972: num 4183 5473 1086 2264 855 ...
$ gdpPercap_1977: num 4910 3009 1029 3215 743 ...
$ gdpPercap_1982: num 5745 2757 1278 4551 807 ...
$ gdpPercap_1987: num 5681 2430 1226 6206 912 ...
$ gdpPercap_1992: num 5023 2628 1191 7954 932 ...
$ gdpPercap_1997: num 4797 2277 1233 8647 946 ...
$ gdpPercap_2002: num 5288 2773 1373 11004 1038 ...
$ gdpPercap_2007: num 6223 4797 1441 12570 1217 ...
$ lifeExp_1952 : num 43.1 30 38.2 47.6 32 ...
$ lifeExp_1957 : num 45.7 32 40.4 49.6 34.9 ...
$ lifeExp_1962 : num 48.3 34 42.6 51.5 37.8 ...
$ lifeExp_1967 : num 51.4 36 44.9 53.3 40.7 ...
$ lifeExp_1972 : num 54.5 37.9 47 56 43.6 ...
$ lifeExp_1977 : num 58 39.5 49.2 59.3 46.1 ...
$ lifeExp_1982 : num 61.4 39.9 50.9 61.5 48.1 ...
$ lifeExp_1987 : num 65.8 39.9 52.3 63.6 49.6 ...
$ lifeExp_1992 : num 67.7 40.6 53.9 62.7 50.3 ...
$ lifeExp_1997 : num 69.2 41 54.8 52.6 50.3 ...
$ lifeExp_2002 : num 71 41 54.4 46.6 50.6 ...
$ lifeExp_2007 : num 72.3 42.7 56.7 50.7 52.3 ...
$ pop_1952 : num 9279525 4232095 1738315 442308 4469979 ...
$ pop_1957 : num 10270856 4561361 1925173 474639 4713416 ...
$ pop_1962 : num 11000948 4826015 2151895 512764 4919632 ...
$ pop_1967 : num 12760499 5247469 2427334 553541 5127935 ...
$ pop_1972 : num 14760787 5894858 2761407 619351 5433886 ...
$ pop_1977 : num 17152804 6162675 3168267 781472 5889574 ...
$ pop_1982 : num 20033753 7016384 3641603 970347 6634596 ...
$ pop_1987 : num 23254956 7874230 4243788 1151184 7586551 ...
$ pop_1992 : num 26298373 8735988 4981671 1342614 8878303 ...
$ pop_1997 : num 29072015 9875024 6066080 1536536 10352843 ...
$ pop_2002 : int 31287142 10866106 7026113 1630347 12251209 7021078 15929988 4048013 8835739 614382 ...
$ pop_2007 : int 33333216 12420476 8078314 1639131 14326203 8390505 17696293 4369038 10238807 710960 ...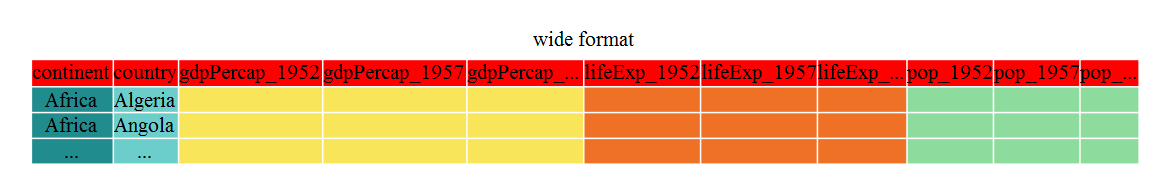
To change this very wide data frame layout back to our nice,
intermediate (or longer) layout, we will use one of the two available
pivot functions from the tidyr package. To
convert from wide to a longer format, we will use the
pivot_longer() function. pivot_longer() makes
datasets longer by increasing the number of rows and decreasing the
number of columns, or ‘lengthening’ your observation variables into a
single variable.
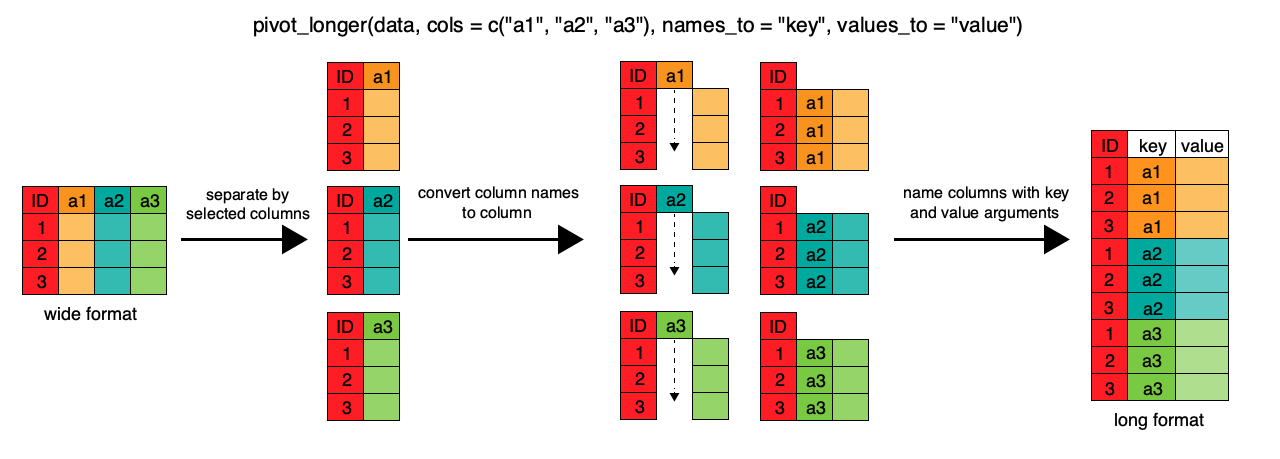
R
gap_long <- gap_wide %>%
pivot_longer(
cols = c(starts_with('pop'), starts_with('lifeExp'), starts_with('gdpPercap')),
names_to = "obstype_year", values_to = "obs_values"
)
str(gap_long)
OUTPUT
tibble [5,112 × 4] (S3: tbl_df/tbl/data.frame)
$ continent : chr [1:5112] "Africa" "Africa" "Africa" "Africa" ...
$ country : chr [1:5112] "Algeria" "Algeria" "Algeria" "Algeria" ...
$ obstype_year: chr [1:5112] "pop_1952" "pop_1957" "pop_1962" "pop_1967" ...
$ obs_values : num [1:5112] 9279525 10270856 11000948 12760499 14760787 ...Here we have used piping syntax which is similar to what we were doing in the previous lesson with dplyr. In fact, these are compatible and you can use a mix of tidyr and dplyr functions by piping them together.
We first provide to pivot_longer() a vector of column
names that will be pivoted into longer format. We could type out all the
observation variables, but as in the select() function (see
dplyr lesson), we can use the starts_with()
argument to select all variables that start with the desired character
string. pivot_longer() also allows the alternative syntax
of using the - symbol to identify which variables are not
to be pivoted (i.e. ID variables).
The next arguments to pivot_longer() are
names_to for naming the column that will contain the new ID
variable (obstype_year) and values_to for
naming the new amalgamated observation variable
(obs_value). We supply these new column names as
strings.
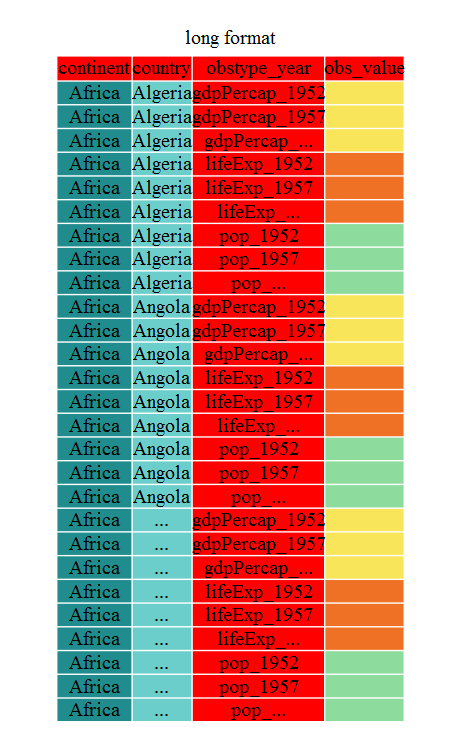
R
gap_long <- gap_wide %>%
pivot_longer(
cols = c(-continent, -country),
names_to = "obstype_year", values_to = "obs_values"
)
str(gap_long)
OUTPUT
tibble [5,112 × 4] (S3: tbl_df/tbl/data.frame)
$ continent : chr [1:5112] "Africa" "Africa" "Africa" "Africa" ...
$ country : chr [1:5112] "Algeria" "Algeria" "Algeria" "Algeria" ...
$ obstype_year: chr [1:5112] "gdpPercap_1952" "gdpPercap_1957" "gdpPercap_1962" "gdpPercap_1967" ...
$ obs_values : num [1:5112] 2449 3014 2551 3247 4183 ...That may seem trivial with this particular data frame, but sometimes you have 1 ID variable and 40 observation variables with irregular variable names. The flexibility is a huge time saver!
Now obstype_year actually contains 2 pieces of
information, the observation type
(pop,lifeExp, or gdpPercap) and
the year. We can use the separate() function
to split the character strings into multiple variables
R
gap_long <- gap_long %>% separate(obstype_year, into = c('obs_type', 'year'), sep = "_")
gap_long$year <- as.integer(gap_long$year)
Challenge 2
Using gap_long, calculate the mean life expectancy,
population, and gdpPercap for each continent. Hint: use
the group_by() and summarize() functions we
learned in the dplyr lesson
R
gap_long %>% group_by(continent, obs_type) %>%
summarize(means=mean(obs_values))
OUTPUT
`summarise()` has regrouped the output.
ℹ Summaries were computed grouped by continent and obs_type.
ℹ Output is grouped by continent.
ℹ Use `summarise(.groups = "drop_last")` to silence this message.
ℹ Use `summarise(.by = c(continent, obs_type))` for per-operation grouping
(`?dplyr::dplyr_by`) instead.OUTPUT
# A tibble: 15 × 3
# Groups: continent [5]
continent obs_type means
<chr> <chr> <dbl>
1 Africa gdpPercap 2194.
2 Africa lifeExp 48.9
3 Africa pop 9916003.
4 Americas gdpPercap 7136.
5 Americas lifeExp 64.7
6 Americas pop 24504795.
7 Asia gdpPercap 7902.
8 Asia lifeExp 60.1
9 Asia pop 77038722.
10 Europe gdpPercap 14469.
11 Europe lifeExp 71.9
12 Europe pop 17169765.
13 Oceania gdpPercap 18622.
14 Oceania lifeExp 74.3
15 Oceania pop 8874672. From long to intermediate format with pivot_wider()
It is always good to check work. So, let’s use the second
pivot function, pivot_wider(), to ‘widen’ our
observation variables back out. pivot_wider() is the
opposite of pivot_longer(), making a dataset wider by
increasing the number of columns and decreasing the number of rows. We
can use pivot_wider() to pivot or reshape our
gap_long to the original intermediate format or the widest
format. Let’s start with the intermediate format.
The pivot_wider() function takes names_from
and values_from arguments.
To names_from we supply the column name whose contents
will be pivoted into new output columns in the widened data frame. The
corresponding values will be added from the column named in the
values_from argument.
R
gap_normal <- gap_long %>%
pivot_wider(names_from = obs_type, values_from = obs_values)
dim(gap_normal)
OUTPUT
[1] 1704 6R
dim(gapminder)
OUTPUT
[1] 1704 6R
names(gap_normal)
OUTPUT
[1] "continent" "country" "year" "gdpPercap" "lifeExp" "pop" R
names(gapminder)
OUTPUT
[1] "country" "year" "pop" "continent" "lifeExp" "gdpPercap"Now we’ve got an intermediate data frame gap_normal with
the same dimensions as the original gapminder, but the
order of the variables is different. Let’s fix that before checking if
they are all.equal().
R
gap_normal <- gap_normal[, names(gapminder)]
all.equal(gap_normal, gapminder)
OUTPUT
[1] "Attributes: < Component \"class\": Lengths (3, 1) differ (string compare on first 1) >"
[2] "Attributes: < Component \"class\": 1 string mismatch >"
[3] "Component \"country\": 1704 string mismatches"
[4] "Component \"pop\": Mean relative difference: 1.634504"
[5] "Component \"continent\": 1212 string mismatches"
[6] "Component \"lifeExp\": Mean relative difference: 0.203822"
[7] "Component \"gdpPercap\": Mean relative difference: 1.162302" R
head(gap_normal)
OUTPUT
# A tibble: 6 × 6
country year pop continent lifeExp gdpPercap
<chr> <int> <dbl> <chr> <dbl> <dbl>
1 Algeria 1952 9279525 Africa 43.1 2449.
2 Algeria 1957 10270856 Africa 45.7 3014.
3 Algeria 1962 11000948 Africa 48.3 2551.
4 Algeria 1967 12760499 Africa 51.4 3247.
5 Algeria 1972 14760787 Africa 54.5 4183.
6 Algeria 1977 17152804 Africa 58.0 4910.R
head(gapminder)
OUTPUT
country year pop continent lifeExp gdpPercap
1 Afghanistan 1952 8425333 Asia 28.801 779.4453
2 Afghanistan 1957 9240934 Asia 30.332 820.8530
3 Afghanistan 1962 10267083 Asia 31.997 853.1007
4 Afghanistan 1967 11537966 Asia 34.020 836.1971
5 Afghanistan 1972 13079460 Asia 36.088 739.9811
6 Afghanistan 1977 14880372 Asia 38.438 786.1134We’re almost there, the original was sorted by country,
then year.
R
gap_normal <- gap_normal %>% arrange(country, year)
all.equal(gap_normal, gapminder)
OUTPUT
[1] "Attributes: < Component \"class\": Lengths (3, 1) differ (string compare on first 1) >"
[2] "Attributes: < Component \"class\": 1 string mismatch >" That’s great! We’ve gone from the longest format back to the intermediate and we didn’t introduce any errors in our code.
Now let’s convert the long all the way back to the wide. In the wide
format, we will keep country and continent as ID variables and pivot the
observations across the 3 metrics
(pop,lifeExp,gdpPercap) and time
(year). First we need to create appropriate labels for all
our new variables (time*metric combinations) and we also need to unify
our ID variables to simplify the process of defining
gap_wide.
R
gap_temp <- gap_long %>% unite(var_ID, continent, country, sep = "_")
str(gap_temp)
OUTPUT
tibble [5,112 × 4] (S3: tbl_df/tbl/data.frame)
$ var_ID : chr [1:5112] "Africa_Algeria" "Africa_Algeria" "Africa_Algeria" "Africa_Algeria" ...
$ obs_type : chr [1:5112] "gdpPercap" "gdpPercap" "gdpPercap" "gdpPercap" ...
$ year : int [1:5112] 1952 1957 1962 1967 1972 1977 1982 1987 1992 1997 ...
$ obs_values: num [1:5112] 2449 3014 2551 3247 4183 ...R
gap_temp <- gap_long %>%
unite(ID_var, continent, country, sep = "_") %>%
unite(var_names, obs_type, year, sep = "_")
str(gap_temp)
OUTPUT
tibble [5,112 × 3] (S3: tbl_df/tbl/data.frame)
$ ID_var : chr [1:5112] "Africa_Algeria" "Africa_Algeria" "Africa_Algeria" "Africa_Algeria" ...
$ var_names : chr [1:5112] "gdpPercap_1952" "gdpPercap_1957" "gdpPercap_1962" "gdpPercap_1967" ...
$ obs_values: num [1:5112] 2449 3014 2551 3247 4183 ...Using unite() we now have a single ID variable which is
a combination of continent,country,and we have
defined variable names. We’re now ready to pipe in
pivot_wider()
R
gap_wide_new <- gap_long %>%
unite(ID_var, continent, country, sep = "_") %>%
unite(var_names, obs_type, year, sep = "_") %>%
pivot_wider(names_from = var_names, values_from = obs_values)
str(gap_wide_new)
OUTPUT
tibble [142 × 37] (S3: tbl_df/tbl/data.frame)
$ ID_var : chr [1:142] "Africa_Algeria" "Africa_Angola" "Africa_Benin" "Africa_Botswana" ...
$ gdpPercap_1952: num [1:142] 2449 3521 1063 851 543 ...
$ gdpPercap_1957: num [1:142] 3014 3828 960 918 617 ...
$ gdpPercap_1962: num [1:142] 2551 4269 949 984 723 ...
$ gdpPercap_1967: num [1:142] 3247 5523 1036 1215 795 ...
$ gdpPercap_1972: num [1:142] 4183 5473 1086 2264 855 ...
$ gdpPercap_1977: num [1:142] 4910 3009 1029 3215 743 ...
$ gdpPercap_1982: num [1:142] 5745 2757 1278 4551 807 ...
$ gdpPercap_1987: num [1:142] 5681 2430 1226 6206 912 ...
$ gdpPercap_1992: num [1:142] 5023 2628 1191 7954 932 ...
$ gdpPercap_1997: num [1:142] 4797 2277 1233 8647 946 ...
$ gdpPercap_2002: num [1:142] 5288 2773 1373 11004 1038 ...
$ gdpPercap_2007: num [1:142] 6223 4797 1441 12570 1217 ...
$ lifeExp_1952 : num [1:142] 43.1 30 38.2 47.6 32 ...
$ lifeExp_1957 : num [1:142] 45.7 32 40.4 49.6 34.9 ...
$ lifeExp_1962 : num [1:142] 48.3 34 42.6 51.5 37.8 ...
$ lifeExp_1967 : num [1:142] 51.4 36 44.9 53.3 40.7 ...
$ lifeExp_1972 : num [1:142] 54.5 37.9 47 56 43.6 ...
$ lifeExp_1977 : num [1:142] 58 39.5 49.2 59.3 46.1 ...
$ lifeExp_1982 : num [1:142] 61.4 39.9 50.9 61.5 48.1 ...
$ lifeExp_1987 : num [1:142] 65.8 39.9 52.3 63.6 49.6 ...
$ lifeExp_1992 : num [1:142] 67.7 40.6 53.9 62.7 50.3 ...
$ lifeExp_1997 : num [1:142] 69.2 41 54.8 52.6 50.3 ...
$ lifeExp_2002 : num [1:142] 71 41 54.4 46.6 50.6 ...
$ lifeExp_2007 : num [1:142] 72.3 42.7 56.7 50.7 52.3 ...
$ pop_1952 : num [1:142] 9279525 4232095 1738315 442308 4469979 ...
$ pop_1957 : num [1:142] 10270856 4561361 1925173 474639 4713416 ...
$ pop_1962 : num [1:142] 11000948 4826015 2151895 512764 4919632 ...
$ pop_1967 : num [1:142] 12760499 5247469 2427334 553541 5127935 ...
$ pop_1972 : num [1:142] 14760787 5894858 2761407 619351 5433886 ...
$ pop_1977 : num [1:142] 17152804 6162675 3168267 781472 5889574 ...
$ pop_1982 : num [1:142] 20033753 7016384 3641603 970347 6634596 ...
$ pop_1987 : num [1:142] 23254956 7874230 4243788 1151184 7586551 ...
$ pop_1992 : num [1:142] 26298373 8735988 4981671 1342614 8878303 ...
$ pop_1997 : num [1:142] 29072015 9875024 6066080 1536536 10352843 ...
$ pop_2002 : num [1:142] 31287142 10866106 7026113 1630347 12251209 ...
$ pop_2007 : num [1:142] 33333216 12420476 8078314 1639131 14326203 ...Challenge 3
Take this 1 step further and create a
gap_ludicrously_wide format data by pivoting over
countries, year and the 3 metrics? Hint this new data
frame should only have 5 rows.
R
gap_ludicrously_wide <- gap_long %>%
unite(var_names, obs_type, year, country, sep = "_") %>%
pivot_wider(names_from = var_names, values_from = obs_values)
Now we have a great ‘wide’ format data frame, but the
ID_var could be more usable, let’s separate it into 2
variables with separate()
R
gap_wide_betterID <- separate(gap_wide_new, ID_var, c("continent", "country"), sep="_")
gap_wide_betterID <- gap_long %>%
unite(ID_var, continent, country, sep = "_") %>%
unite(var_names, obs_type, year, sep = "_") %>%
pivot_wider(names_from = var_names, values_from = obs_values) %>%
separate(ID_var, c("continent","country"), sep = "_")
str(gap_wide_betterID)
OUTPUT
tibble [142 × 38] (S3: tbl_df/tbl/data.frame)
$ continent : chr [1:142] "Africa" "Africa" "Africa" "Africa" ...
$ country : chr [1:142] "Algeria" "Angola" "Benin" "Botswana" ...
$ gdpPercap_1952: num [1:142] 2449 3521 1063 851 543 ...
$ gdpPercap_1957: num [1:142] 3014 3828 960 918 617 ...
$ gdpPercap_1962: num [1:142] 2551 4269 949 984 723 ...
$ gdpPercap_1967: num [1:142] 3247 5523 1036 1215 795 ...
$ gdpPercap_1972: num [1:142] 4183 5473 1086 2264 855 ...
$ gdpPercap_1977: num [1:142] 4910 3009 1029 3215 743 ...
$ gdpPercap_1982: num [1:142] 5745 2757 1278 4551 807 ...
$ gdpPercap_1987: num [1:142] 5681 2430 1226 6206 912 ...
$ gdpPercap_1992: num [1:142] 5023 2628 1191 7954 932 ...
$ gdpPercap_1997: num [1:142] 4797 2277 1233 8647 946 ...
$ gdpPercap_2002: num [1:142] 5288 2773 1373 11004 1038 ...
$ gdpPercap_2007: num [1:142] 6223 4797 1441 12570 1217 ...
$ lifeExp_1952 : num [1:142] 43.1 30 38.2 47.6 32 ...
$ lifeExp_1957 : num [1:142] 45.7 32 40.4 49.6 34.9 ...
$ lifeExp_1962 : num [1:142] 48.3 34 42.6 51.5 37.8 ...
$ lifeExp_1967 : num [1:142] 51.4 36 44.9 53.3 40.7 ...
$ lifeExp_1972 : num [1:142] 54.5 37.9 47 56 43.6 ...
$ lifeExp_1977 : num [1:142] 58 39.5 49.2 59.3 46.1 ...
$ lifeExp_1982 : num [1:142] 61.4 39.9 50.9 61.5 48.1 ...
$ lifeExp_1987 : num [1:142] 65.8 39.9 52.3 63.6 49.6 ...
$ lifeExp_1992 : num [1:142] 67.7 40.6 53.9 62.7 50.3 ...
$ lifeExp_1997 : num [1:142] 69.2 41 54.8 52.6 50.3 ...
$ lifeExp_2002 : num [1:142] 71 41 54.4 46.6 50.6 ...
$ lifeExp_2007 : num [1:142] 72.3 42.7 56.7 50.7 52.3 ...
$ pop_1952 : num [1:142] 9279525 4232095 1738315 442308 4469979 ...
$ pop_1957 : num [1:142] 10270856 4561361 1925173 474639 4713416 ...
$ pop_1962 : num [1:142] 11000948 4826015 2151895 512764 4919632 ...
$ pop_1967 : num [1:142] 12760499 5247469 2427334 553541 5127935 ...
$ pop_1972 : num [1:142] 14760787 5894858 2761407 619351 5433886 ...
$ pop_1977 : num [1:142] 17152804 6162675 3168267 781472 5889574 ...
$ pop_1982 : num [1:142] 20033753 7016384 3641603 970347 6634596 ...
$ pop_1987 : num [1:142] 23254956 7874230 4243788 1151184 7586551 ...
$ pop_1992 : num [1:142] 26298373 8735988 4981671 1342614 8878303 ...
$ pop_1997 : num [1:142] 29072015 9875024 6066080 1536536 10352843 ...
$ pop_2002 : num [1:142] 31287142 10866106 7026113 1630347 12251209 ...
$ pop_2007 : num [1:142] 33333216 12420476 8078314 1639131 14326203 ...R
all.equal(gap_wide, gap_wide_betterID)
OUTPUT
[1] "Attributes: < Component \"class\": Lengths (1, 3) differ (string compare on first 1) >"
[2] "Attributes: < Component \"class\": 1 string mismatch >" There and back again!
Other great resources
- R for Data Science (online book)
- Data Wrangling Cheat sheet (pdf file)
- Introduction to tidyr (online documentation)
- Data wrangling with R and RStudio (online video)
- Use the
tidyrpackage to change the layout of data frames. - Use
pivot_longer()to go from wide to longer layout. - Use
pivot_wider()to go from long to wider layout.
Content from Producing Reports With knitr
Last updated on 2026-04-21 | Edit this page
Estimated time: 75 minutes
Overview
Questions
- How can I integrate software and reports?
Objectives
- Understand the value of writing reproducible reports
- Learn how to recognise and compile the basic components of an R Markdown file
- Become familiar with R code chunks, and understand their purpose, structure and options
- Demonstrate the use of inline chunks for weaving R outputs into text blocks, for example when discussing the results of some calculations
- Be aware of alternative output formats to which an R Markdown file can be exported
Data analysis reports
Data analysts tend to write a lot of reports, describing their analyses and results, for their collaborators or to document their work for future reference.
Many new users begin by first writing a single R script containing all of their work, and then share the analysis by emailing the script and various graphs as attachments. But this can be cumbersome, requiring a lengthy discussion to explain which attachment was which result.
Writing formal reports with Word or LaTeX can simplify this process by incorporating both the analysis report and output graphs into a single document. But tweaking formatting to make figures look correct and fixing obnoxious page breaks can be tedious and lead to a lengthy “whack-a-mole” game of fixing new mistakes resulting from a single formatting change.
Creating a report as a web page (which is an html file) using R Markdown makes things easier. The report can be one long stream, so tall figures that wouldn’t ordinarily fit on one page can be kept at full size and easier to read, since the reader can simply keep scrolling. Additionally, the formatting of and R Markdown document is simple and easy to modify, allowing you to spend more time on your analyses instead of writing reports.
Literate programming
Ideally, such analysis reports are reproducible documents: If an error is discovered, or if some additional subjects are added to the data, you can re-compile the report and get the new or corrected results rather than having to reconstruct figures, paste them into a Word document, and hand-edit various detailed results.
The key R package here is knitr. It allows you
to create a document that is a mixture of text and chunks of code. When
the document is processed by knitr, chunks of code will be
executed, and graphs or other results will be inserted into the final
document.
This sort of idea has been called “literate programming”.
knitr allows you to mix basically any type of text with
code from different programming languages, but we recommend that you use
R Markdown, which mixes Markdown with R. Markdown is a light-weight
mark-up language for creating web pages.
Creating an R Markdown file
Within RStudio, click File → New File → R Markdown and you’ll get a dialog box like this:
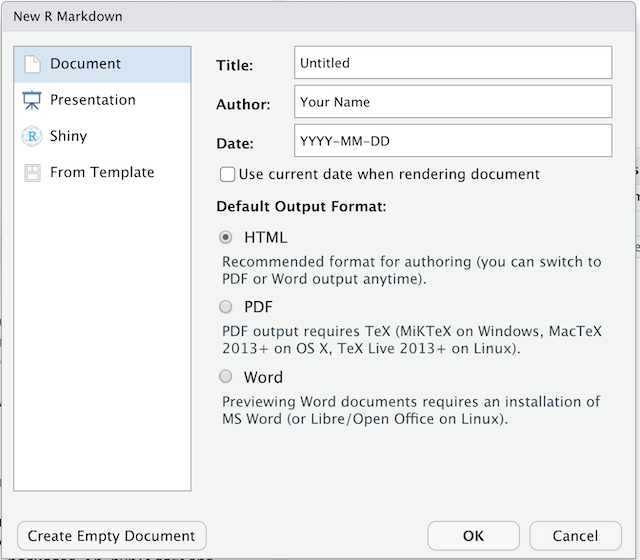
You can stick with the default (HTML output), but give it a title.
Basic components of R Markdown
The initial chunk of text (header) contains instructions for R to specify what kind of document will be created, and the options chosen. You can use the header to give your document a title, author, date, and tell it what type of output you want to produce. In this case, we’re creating an html document.
---
title: "Initial R Markdown document"
author: "Karl Broman"
date: "April 23, 2015"
output: html_document
---You can delete any of those fields if you don’t want them included. The double-quotes aren’t strictly necessary in this case. They’re mostly needed if you want to include a colon in the title.
RStudio creates the document with some example text to get you started. Note below that there are chunks like
```{r}
summary(cars)
```
These are chunks of R code that will be executed by
knitr and replaced by their results. More on this
later.
Markdown
Markdown is a system for writing web pages by marking up the text much as you would in an email rather than writing html code. The marked-up text gets converted to html, replacing the marks with the proper html code.
For now, let’s delete all of the stuff that’s there and write a bit of markdown.
You make things bold using two asterisks, like this:
**bold**, and you make things italics by using
underscores, like this: _italics_.
You can make a bulleted list by writing a list with hyphens or asterisks with a space between the list and other text, like this:
A list:
* bold with double-asterisks
* italics with underscores
* code-type font with backticksor like this:
A second list:
- bold with double-asterisks
- italics with underscores
- code-type font with backticksEach will appear as:
- bold with double-asterisks
- italics with underscores
- code-type font with backticks
You can use whatever method you prefer, but be consistent. This maintains the readability of your code.
You can make a numbered list by using numbers instead. You can even use the same number over and over if you want:
1. bold with double-asterisks
1. italics with underscores
1. code-type font with backticksThis will appear as:
- bold with double-asterisks
- italics with underscores
- code-type font with backticks
You can make section headers of different sizes by initiating a line
with some number of # symbols:
# Title
## Main section
### Sub-section
#### Sub-sub sectionYou compile the R Markdown document to an html webpage by clicking the “Knit” button in the upper-left.
Challenge 1
Create a new R Markdown document. Delete all of the R code chunks and write a bit of Markdown (some sections, some italicized text, and an itemized list).
Convert the document to a webpage.
In RStudio, select File > New file > R Markdown…
Delete the placeholder text and add the following:
# Introduction
## Background on Data
This report uses the *gapminder* dataset, which has columns that include:
* country
* continent
* year
* lifeExp
* pop
* gdpPercap
## Background on Methods
Then click the ‘Knit’ button on the toolbar to generate an html document (webpage).
A bit more Markdown
You can make a hyperlink like this:
[Carpentries Home Page](https://carpentries.org/).
You can include an image file like this:

You can do subscripts (e.g., F2) with F~2~
and superscripts (e.g., F2) with F^2^.
If you know how to write equations in LaTeX, you can use
$ $ and $$ $$ to insert math equations, like
$E = mc^2$ and
$$y = \mu + \sum_{i=1}^p \beta_i x_i + \epsilon$$You can review Markdown syntax by navigating to the “Markdown Quick Reference” under the “Help” field in the toolbar at the top of RStudio.
R code chunks
The real power of Markdown comes from mixing markdown with chunks of code. This is R Markdown. When processed, the R code will be executed; if they produce figures, the figures will be inserted in the final document.
The main code chunks look like this:
```{r load_data}
gapminder <- read.csv("gapminder.csv")
```
That is, you place a chunk of R code between ```{r
chunk_name} and ```. You should give each chunk a
unique name, as they will help you to fix errors and, if any graphs are
produced, the file names are based on the name of the code chunk that
produced them. You can create code chunks quickly in RStudio using the
shortcuts Ctrl+Alt+I on Windows and
Linux, or Cmd+Option+I on Mac.
Challenge 2
Add code chunks to:
- Load the ggplot2 package
- Read the gapminder data
- Create a plot
```{r load-ggplot2}
library("ggplot2")
```
```{r read-gapminder-data}
gapminder <- read.csv("gapminder.csv")
```
```{r make-plot}
plot(lifeExp ~ year, data = gapminder)
```
How things get compiled
When you press the “Knit” button, the R Markdown document is
processed by knitr
and a plain Markdown document is produced (as well as, potentially, a
set of figure files): the R code is executed and replaced by both the
input and the output; if figures are produced, links to those figures
are included.
The Markdown and figure documents are then processed by the tool pandoc, which converts the
Markdown file into an html file, with the figures embedded.
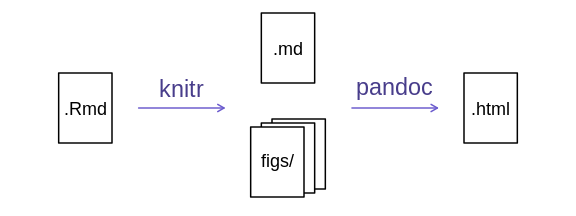
Chunk options
There are a variety of options to affect how the code chunks are treated. Here are some examples:
- Use
echo=FALSEto avoid having the code itself shown. - Use
results="hide"to avoid having any results printed. - Use
eval=FALSEto have the code shown but not evaluated. - Use
warning=FALSEandmessage=FALSEto hide any warnings or messages produced. - Use
fig.heightandfig.widthto control the size of the figures produced (in inches).
So you might write:
```{r load_libraries, echo=FALSE, message=FALSE}
library("dplyr")
library("ggplot2")
```
Often there will be particular options that you’ll want to use repeatedly; for this, you can set global chunk options, like so:
```{r global_options, echo=FALSE}
knitr::opts_chunk$set(fig.path="Figs/", message=FALSE, warning=FALSE,
echo=FALSE, results="hide", fig.width=11)
```
The fig.path option defines where the figures will be
saved. The / here is really important; without it, the
figures would be saved in the standard place but just with names that
begin with Figs.
If you have multiple R Markdown files in a common directory, you
might want to use fig.path to define separate prefixes for
the figure file names, like fig.path="Figs/cleaning-" and
fig.path="Figs/analysis-".
Challenge 3
Use chunk options to control the size of a figure and to hide the code.
```{r echo = FALSE, fig.width = 3}
plot(faithful)
```
You can review all of the R chunk options by navigating
to the “R Markdown Cheat Sheet” under the “Cheatsheets” section of the
“Help” field in the toolbar at the top of RStudio.
Inline R code
You can make every number in your report reproducible. Use
`r and ` for an in-line code chunk, like so:
`r round(some_value, 2)`. The code will be executed and
replaced with the value of the result.
Don’t let these in-line chunks get split across lines.
Perhaps precede the paragraph with a larger code chunk that does
calculations and defines variables, with include=FALSE for
that larger chunk (which is the same as echo=FALSE and
results="hide").
Rounding can produce differences in output in such situations. You
may want 2.0, but round(2.03, 1) will give
just 2.
The myround
function in the R/broman
package handles this.
Challenge 4
Try out a bit of in-line R code.
Here’s some inline code to determine that 2 + 2 = 4.
Other output options
You can also convert R Markdown to a PDF or a Word document. Click
the little triangle next to the “Knit” button to get a drop-down menu.
Or you could put pdf_document or word_document
in the initial header of the file.
Tip: Creating PDF documents
Creating .pdf documents may require installation of some extra
software. The R package tinytex provides some tools to help
make this process easier for R users. With tinytex
installed, run tinytex::install_tinytex() to install the
required software (you’ll only need to do this once) and then when you
knit to pdf tinytex will automatically detect and install
any additional LaTeX packages that are needed to produce the pdf
document. Visit the tinytex
website for more information.
Tip: Visual markdown editing in RStudio
RStudio versions 1.4 and later include visual markdown editing mode.
In visual editing mode, markdown expressions (like
**bold words**) are transformed to the formatted appearance
(bold words) as you type. This mode also includes a
toolbar at the top with basic formatting buttons, similar to what you
might see in common word processing software programs. You can turn
visual editing on and off by pressing the ![]() button in the top right corner of your R Markdown document.
button in the top right corner of your R Markdown document.
Resources
- Knitr in a knutshell tutorial
- Dynamic Documents with R and knitr (book)
- R Markdown documentation
- R Markdown cheat sheet
- Getting started with R Markdown
- R Markdown: The Definitive Guide (book by Rstudio team)
- Reproducible Reporting
- The Ecosystem of R Markdown
- Introducing Bookdown
- Mix reporting written in R Markdown with software written in R.
- Specify chunk options to control formatting.
- Use
knitrto convert these documents into PDF and other formats.
Content from Writing Good Software
Last updated on 2026-04-21 | Edit this page
Estimated time: 15 minutes
Overview
Questions
- How can I write software that other people can use?
Objectives
- Describe best practices for writing R and explain the justification for each.
Structure your project folder
Keep your project folder structured, organized and tidy, by creating
subfolders for your code files, manuals, data, binaries, output plots,
etc. It can be done completely manually, or with the help of RStudio’s
New Project functionality, or a designated package, such as
ProjectTemplate.
Tip: ProjectTemplate - a possible solution
One way to automate the management of projects is to install the
third-party package, ProjectTemplate. This package will set
up an ideal directory structure for project management. This is very
useful as it enables you to have your analysis pipeline/workflow
organised and structured. Together with the default RStudio project
functionality and Git you will be able to keep track of your work as
well as be able to share your work with collaborators.
- Install
ProjectTemplate. - Load the library
- Initialise the project:
R
install.packages("ProjectTemplate")
library("ProjectTemplate")
create.project("../my_project_2", merge.strategy = "allow.non.conflict")
For more information on ProjectTemplate and its functionality visit the home page ProjectTemplate
Make code readable
The most important part of writing code is making it readable and understandable. You want someone else to be able to pick up your code and be able to understand what it does: more often than not this someone will be you 6 months down the line, who will otherwise be cursing past-self.
Documentation: tell us what and why, not how
When you first start out, your comments will often describe what a command does, since you’re still learning yourself and it can help to clarify concepts and remind you later. However, these comments aren’t particularly useful later on when you don’t remember what problem your code is trying to solve. Try to also include comments that tell you why you’re solving a problem, and what problem that is. The how can come after that: it’s an implementation detail you ideally shouldn’t have to worry about.
Keep your code modular
Our recommendation is that you should separate your functions from
your analysis scripts, and store them in a separate file that you
source when you open the R session in your project. This
approach is nice because it leaves you with an uncluttered analysis
script, and a repository of useful functions that can be loaded into any
analysis script in your project. It also lets you group related
functions together easily.
Break down problem into bite size pieces
When you first start out, problem solving and function writing can be daunting tasks, and hard to separate from code inexperience. Try to break down your problem into digestible chunks and worry about the implementation details later: keep breaking down the problem into smaller and smaller functions until you reach a point where you can code a solution, and build back up from there.
Know that your code is doing the right thing
Make sure to test your functions!
Don’t repeat yourself
Functions enable easy reuse within a project. If you see blocks of similar lines of code through your project, those are usually candidates for being moved into functions.
If your calculations are performed through a series of functions, then the project becomes more modular and easier to change. This is especially the case for which a particular input always gives a particular output.
Remember to be stylish
Apply consistent style to your code.
- Keep your project folder structured, organized and tidy.
- Document what and why, not how.
- Break programs into short single-purpose functions.
- Write re-runnable tests.
- Don’t repeat yourself.
- Be consistent in naming, indentation, and other aspects of style.
